Volume 10, Number 9—September 2004
Dispatch
Typing of Borrelia Relapsing Fever Group Strains
Abstract
Partial sequencing of the 16S-23S rDNA intergenic spacer showed two to four genotypes each for Borrelia hermsii and B. turicatae, both relapsing fever agents transmitted by argasid ticks, and for B. miyamotoi and B. lonestari, transmitted by ixodid ticks. Field surveys of Ixodes ticks in Connecticut and Sweden showed limited local diversity for B. miyamotoi.
The two major clades of species in the genus Borrelia are the Lyme borreliosis group and the relapsing fever group (1). The Lyme borreliosis group includes Borrelia burgdorferi, B. afzelii, B. garinii, and several other species not associated with human disease. The relapsing fever group includes several species, such as B. hermsii in the Nearctic ecologic region and B. persica in the Palearctic, that cause endemic relapsing fever in humans (2). The known relapsing fever agents are transmitted by soft (argasid) ticks, usually an Ornithodoros species. In 1995, B. miyamotoi was first isolated from Ixodes persulcatus hard (ixodid) ticks in Japan (3). Genomic DNA of the newly identified spirochete cross-hybridized to a greater extent with DNA of relapsing fever species than with DNA of Lyme borreliosis species. In 1996, B. lonestari was discovered in Amblyomma americanum, an ixodid tick of the southern and eastern United States (4,5). Although B. lonestari is associated with a Lyme borreliosis–like disorder in the southern United States (6), sequence analysis showed that B. lonestari, like B. miyamotoi, was in a clade with the relapsing fever group rather than the Lyme borreliosis group (4,5). More recently, spirochetes closely related to B. miyamotoi, and provisionally designated here as B. miyamotoi sensu lato (s.l.), were discovered in I. scapularis ticks in the United States (7) and I. ricinus ticks in Europe (8).
The public health importance of the newly discovered species remains to be determined. However, finding B. miyamotoi s.l. in I. scapularis, I. ricinus, and I. persulcatus, the predominant vectors of Lyme borreliosis in North America, Europe, and Asia, respectively, complicates interpreting epidemiologic studies of Lyme borreliosis and other ixodid-borne disorders. A method to identify and distinguish strains within species is needed to carry out studies of the population biology and of the possible etiologic roles of these organisms. Since most of these microorganisms are to date uncultivable or poorly cultivable, a method using DNA amplification by polymerase chain reaction (PCR) is preferable. On the basis of the findings of Liveris et al. (9), we further developed sequence analysis of the 16S-23S rRNA intergenic spacer (IGS) for strain typing and showed its advantages over other loci for the Lyme borreliosis agents B. burgdorferi and B. afzelii (10). For this study, we applied this approach to typing the new Borrelia spp. and included two relapsing fever agents, B. hermsii (endemic in the western and northwestern United States) and B. turicatae (endemic in the southwestern and south-central United States) (2).
Nine isolates of B. hermsii in our culture collection originated in New Mexico, Colorado, California, and Washington State and were either from Ornithodoros hermsi ticks, patients with relapsing fever, or, in one case, a bird (11). Two B. turicatae isolates were from O. turicata ticks from Texas and Kansas. B. miyamotoi strains HT24, HT31, and HK004 from I. persulcatus ticks and strains NB103-1 and FR64b from Apodemus spp. mice were from Hokkaido, Japan (3,12). Cultivable strains of these species were grown in Barbour-Stoenner-Kelly II medium. Uncultivated species were initially identified in total DNA extracts of ticks by using Borrelia genus–specific PCR, targeting flaB gene (5). Approximately 2% of A. americanum nymphs and adult females in collections from different areas of New Jersey, Illinois, and Missouri contained B. lonestari. B. miyamotoi s.l. spirochetes were identified in I. scapularis nymphs collected at a 7.2-ha field site in southern Connecticut and in I. ricinus nymphs collected at a 1.5-ha site in Blekinge County in Sweden (10). A Connecticut strain of B. miyamotoi s.l. strain was maintained in Mus musculus (7).
Part of the intergenic spacer was amplified by PCR with primers for the 3′ end of the 16S rRNA gene and the ileT tRNA gene (10). As a comparison to the intergenic spacer locus and to assess linkage disequilibrium, we also partially sequenced the chromosomal gene for the P66 outer membrane protein (10,13) after amplification by PCR as described in the Table footnotes. The PCR products were either directly sequenced or first cloned into pCR2.1-TOPO vector (Invitrogen, Carlsbad, CA) before sequencing on a Beckman CEQ 8000 (Beckman Coulter, Fullerton, CA) automated sequencer. The sequences were aligned automatically by using Clustal X software (http://www-igbmc.u-strasbg.fr/BioInfo/ClustalX) and then manually with MacCLADE version 4.05 (http://macclade.org/macclade.html) (10). The maximum lengths of the alignments (http://spiro.mmg.uci.edu/data) were set by the shortest available sequence. Accession numbers for the deposited sequences are given in the legend for the Figure and in a footnote for the Table.
The PCR products for the intergenic spacer locus varied in length between species and ranged from 388 bp for B. miyamotoi s.l. from Sweden to 685 bp for B. turicatae. The PCR product for the p66 gene was 605–614 bp between species. The Table summarizes the statistics for the aligned intergenic spacer and p66 sequences of the B. miyamotoi s.l., B. lonestari, and B. hermsii. The mean nucleotide diversity normalized for each aligned position was 38%-130% higher for the intergenic spacer locus than for the p66 locus. At the same time, intragenic recombination was not detected at the intergenic spacer locus with Sawyer’s test (www.math.wustl.edu/~sawyer/mbprogs), which assesses the likelihood that polymorphisms in a sequence arose through recombination rather than mutation (data not shown). This result was consistent with the undetectable recombination at the intergenic spacer loci of B. burgdorferi (10).
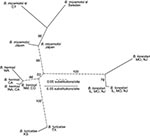
The genetic diversity at the intergenic spacer and p66 loci for the relapsing fever group species in a given geographic area was more limited than was the case for Lyme borreliosis species (10). This limitation was most apparent with the B. miyamotoi s.l. sequences of 22 samples from Connecticut and 6 samples from Sweden. As shown by the phylogram (Figure), only one intergenic spacer genotype each was found for B. miyamotoi s.l. from the Connecticut site and from Sweden. In contrast, collections at the same sites and times, and from the same tick vectors, provided 8 intergenic spacer genotypes among 62 B. burgdorferi samples in I. scapularis and 9 intergenic spacer genotypes among 73 B. afzelii samples in I. ricinus (10). Accepting a type I error level of 0.05, we would expect to have detected a second genotype of B. miyamotoi s.l. in a sample size of 22 if its proportion was >14%. The findings at the p66 locus for 10 Connecticut samples and for 4 samples from Sweden were similar: only one p66 genotype was detected at each location.
The samples of the other relapsing fever group species were not prospectively acquired for population studies, and thus, the findings provide only a tentative view of population structure. Nevertheless, the results are consistent with an interpretation that the local strain diversity of the relapsing fever group species is more limited than that of Lyme borreliosis agents. The intergenic spacer sequences of five B. miyamotoi isolates from ticks or mice from Japan were identical, except for a single nucleotide in one isolate (Figure); the p66 sequences were identical for each of the five isolates. Four intergenic spacer genotypes were detected from the nine isolates of B. hermsii from different regions of the western United States; the three intergenic spacer genotypes that were examined each had a different p66 allele. Two of the linked intergenic spacer and p66 genotypes were unique to species from the Rocky Mountain region. The two strains of B. turicatae from Texas and Kansas differed in intergenic spacer genotype. A. americanum ticks collected in three states yielded three intergenic spacer genotypes from 20 samples positive for B. lonestari (Table and Figure). The three intergenic spacer genotypes were each linked to three unique p66 alleles. Two of the linked genotypes were found at all three locations; one was found in Missouri and New Jersey but not in Illinois.
Samples of the B. miyamotoi s.l. showed greater genetic diversity at the intergenic spacer locus in than did samples from other genomic groups (Figure). However, even for species with fewer polymorphisms (Table), the intergenic spacer sequences, with or without the p66 sequences, confirmed the monophyly of strains within each species. This pattern of relationship and the lack of evidence of gene conversion from horizontal gene transfer at this locus, demonstrates that, as for the Lyme borreliosis spirochetes (10), the intergenic spacer region is both sensitive and sufficient for genotyping the relapsing fever group of Borrelia species. Sequencing this locus provides a means for further epidemiologic and ecologic studies of the newly discovered Borrelia species of hard ticks, as well as of the relapsing fever agents that are reemerging as human pathogens (1). Certain intergenic spacer genotypes of B. burgdorferi are associated with certain virulence phenotypes in humans (14). Strain typing by PCR and sequence analysis should also be useful for identifying and characterizing the vertebrate reservoirs of B. lonestari and B. miyamotoi s.l.
The linkage disequilibrium between the intergenic spacer and p66 loci indicate that the relapsing fever group species, like Lyme borreliosis spirochetes (10,15), are highly clonal bacteria. Why these two groups of tick-borne spirochetes appear to have different population structures remains to be determined. This variation may be the consequence of differences in pathogenesis between the organisms.
Dr. Bunikis is a faculty member in the Department of Microbiology and Molecular Genetics, University of California—Irvine. His research interests include strategies for molecular typing of Borrelia spirochetes for better understanding of their diversity, evolution, ecology, and epidemiologic importance.
Acknowledgments
We thank H. Mirian and H. Mattaous for technical assistance; M. Fukunaga, J. Piesman, B. Schneider, M. Quintana, T. Schulze, T. Walker, and R. Hu for providing Borrelia strains and tick specimens; I. Eliasson for support; and the Southern Sweden Borrelia Study Group for assisting with tick collection.
This work was funded by grants to J. Bunikis (919558-01 from the Centers for Disease Control and Prevention), J. Tsao (National Science Foundation), J. Berglund (Medical faculty of Lund University and the County Council of Blekinge), D. Fish (The Harold G. and Leila Y. Mathers Charitable Foundation and 58-1265-5023 from the United States Department of Agriculture), and A. Barbour (AI37248 from the National Institutes of Health).
References
- Barbour AG. Borrelia: a diverse and ubiquitous genus of tick-borne pathogens. In: Scheld WM, Craig WA, Hughes JM, editors. Emerging infections 5. Washington: ASM Press; 2001. p. 153–74.
- Barbour AG. Specificity of Borrelia-tick vector relationships. SGM Symposium 63: Microbe-vector interactions in vector-borne disease. Cambridge: Cambridge University Press; 2004. p. 75–90.
- Fukunaga M, Takahashi Y, Tsuruta Y, Matsushita O, Ralph D, McClelland M, Genetic and phenotypic analysis of Borrelia miyamotoi sp. nov., isolated from the ixodid tick Ixodes persulcatus, the vector for Lyme disease in Japan. Int J Syst Bacteriol. 1995;45:804–10. DOIPubMedGoogle Scholar
- Armstrong PM, Rich SM, Smith RD, Hartl DL, Spielman A, Telford SR III. A new Borrelia infecting Lone Star ticks. Lancet. 1996;347:67–8. DOIPubMedGoogle Scholar
- Barbour AG, Maupin GO, Teltow GJ, Carter CJ, Piesman J. Identification of an uncultivable Borrelia species in the hard tick Amblyomma americanum: possible agent of a Lyme disease–like illness. J Infect Dis. 1996;173:403–9.PubMedGoogle Scholar
- James AM, Liveris D, Wormser GP, Schwartz I, Montecalvo MA, Johnson BJ. Borrelia lonestari infection after a bite by an Amblyomma americanum tick. J Infect Dis. 2001;183:1810–4. DOIPubMedGoogle Scholar
- Scoles GA, Papero M, Beati L, Fish D. A relapsing fever group spirochete transmitted by Ixodes scapularis ticks. Vector Borne Zoonotic Dis. 2001;1:21–34. DOIPubMedGoogle Scholar
- Fraenkel CJ, Garpmo U, Berglund J. Determination of novel Borrelia genospecies in Swedish Ixodes ricinus ticks. J Clin Microbiol. 2002;40:3308–12. DOIPubMedGoogle Scholar
- Liveris D, Gazumyan A, Schwartz I. Molecular typing of Borrelia burgdorferi sensu lato by PCR-restriction fragment length polymorphism analysis. J Clin Microbiol. 1995;33:589–95.PubMedGoogle Scholar
- Bunikis J, Garpmo U, Tsao J, Berglund J, Fish D, Barbour AG. Sequence typing reveals extensive strain diversity of the Lyme borreliosis agents Borrelia burgdorferi in North America and Borrelia afzelii in Europe. Microbiology. 2004;150:1741–55. DOIPubMedGoogle Scholar
- Thomas NJ, Bunikis J, Barbour AG, Wolcott MJ. Fatal spirochetosis due to a relapsing fever-like Borrelia sp. in a northern spotted owl. J Wildl Dis. 2002;38:187–93.PubMedGoogle Scholar
- Hamase A, Takahashi Y, Nohgi K, Fukunaga M. Homology of variable major protein genes between Borrelia hermsii and Borrelia miyamotoi. FEMS Microbiol Lett. 1996;140:131–7. DOIPubMedGoogle Scholar
- Bunikis J, Luke CJ, Bunikiene E, Bergstrom S, Barbour AG. A surface-exposed region of a novel outer membrane protein (P66) of Borrelia spp. is variable in size and sequence. J Bacteriol. 1998;180:1618–23.PubMedGoogle Scholar
- Wormser GP, Liveris D, Nowakowski J, Nadelman RB, Cavaliere LF, McKenna D, Association of specific subtypes of Borrelia burgdorferi with hematogenous dissemination in early Lyme disease. J Infect Dis. 1999;180:720–5. DOIPubMedGoogle Scholar
- Dykhuizen DE, Polin DS, Dunn JJ, Wilske B, Preac-Mursic V, Dattwyler RJ, Borrelia burgdorferi is clonal: implications for taxonomy and vaccine development. Proc Natl Acad Sci U S A. 1993;90:10163–7. DOIPubMedGoogle Scholar
Figure
Table
Cite This ArticleTable of Contents – Volume 10, Number 9—September 2004
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Jonas Bunikis, Department of Microbiology and Molecular Genetics, B240 Medical Sciences I, University of California—Irvine, Irvine, CA 92697-4025, USA; fax 949-824-6452
Top