Volume 11, Number 1—January 2005
Dispatch
A Novel Paramyxovirus?
Figure 2
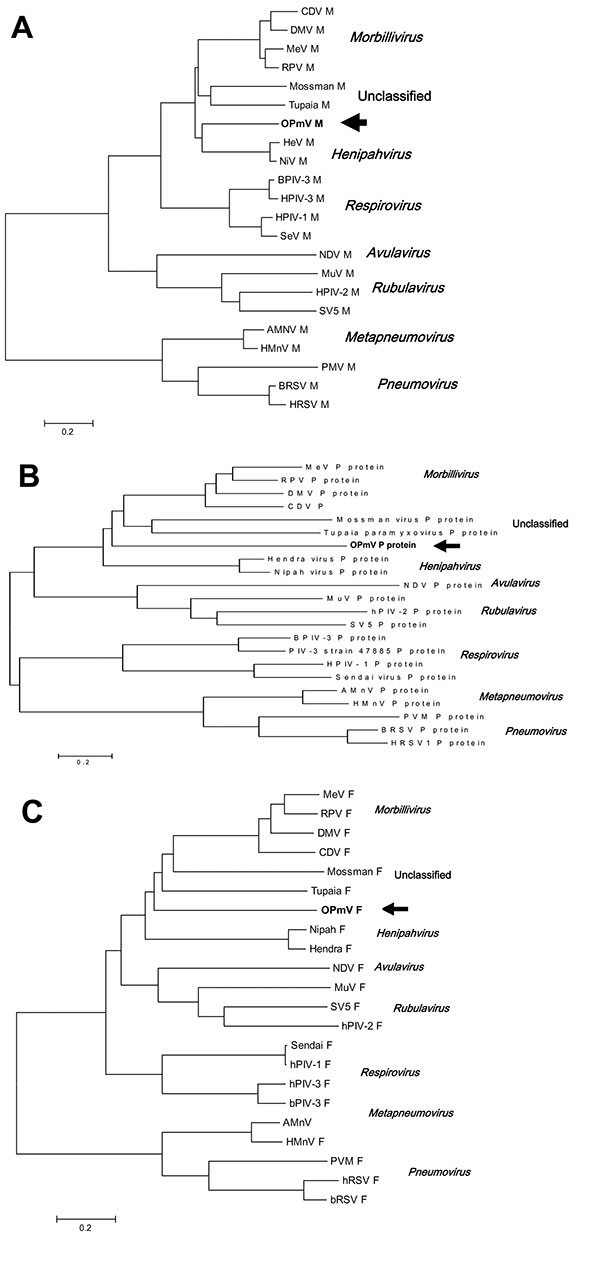
Figure 2. Phylogenetic comparison of OPmV proteins to other paramyxovirus proteins. A) Phylogenetic tree showing the relationship of the putative OPmV M protein to the M proteins of other paramyxoviruses representative of the various genera in the family Paramyxoviridae. B) Phylogenetic tree showing the relationship of the putative OPmV F protein to the F proteins of other representative paramyxoviruses. C) Phylogenetic tree showing the relationship of the putative OPmV P protein to the P proteins of other representative paramyxoviruses. Sequence alignments were made with the ClustalW method of the AlignX program of the Vector NTI6 software package. The trees were generated from these alignments by using neighbor-joining methods through the computer program MEGA version 2.1 (available from http://www.megasoftware.net/). The position of the putative OPmV sequences are indicated by arrows; distance bars, which represent 0.2 amino acid changes per position, are shown below the trees. The sequences from which the trees were constructed are as follows: Mossman, Mossman virus (NC_005339); Tupaia, Tupaia paramyxovirus (NC_002199); NiV, Nipah virus (NC_002728); HeV, Hendra virus (NC_001906); SeV, Sendai virus (AB065188); hPIV-1, human parainfluenza virus 1 (NC_003461); hPIV-3, human parainfluenza virus type 3 (NC_001796); bPIV-3, bovine parainfluenza virus type 3 (AF178655); hRSV, human respiratory syncytial virus (GI:133665); BRSV, Bovine respiratory syncytial virus (NC_001989); PMV, Pneumonia virus of mice (AY573814)AMnV, avian metapneumovirus (AY028582); HMnV, human metapneumovirus (NC_004148); SV5, simian paramyxovirus SV5 (D13868); hPIV-2, human parainfluenza virus type 2 (NC_003443); MuV, mumps virus (AY309060); NDV, Newcastle disease virus (NC_002617); MeV, measles virus (AF266288); RPV, rinderpest virus (AB021977, M21514, M34018); DMV, dolphin morbillivirus (NC_005283); CDV, canine distemper virus (NC_005283).