Volume 11, Number 3—March 2005
Dispatch
Methicillin-resistant Staphylococcus aureus in Neonatal Intensive Care Unit
Abstract
A neonatal intensive care unit outbreak was caused by a strain of methicillin-resistant Staphylococcus aureus previously found in the community (ST45-MRSA-IV). Fifteen infected neonates were identified, 2 of whom died. This outbreak illustrates how a rare community pathogen can rapidly spread through nosocomial transmission.
Since 1998, strains of highly virulent, community-associated, methicillin-resistant Staphylococcus aureus (CA-MRSA), which are distinct from the typical nosocomial MRSA (NA-MRSA), have been reported (1,2). CA-MRSA is susceptible to numerous antimicrobial agents, in contrast to the multidrug-resistant (MDR) NA-MRSA phenotype, because it carries the staphylococcal cassette chromosome mec (SCCmec) type IV or V, rather than type I or III (3,4). The high virulence of CA-MRSA has been linked to Panton-Valentine leukocidin (PVL), a virulence factor found in most of these strains (2,5). We report a nosocomial MRSA outbreak in a neonatal intensive-care unit (NICU), by a non-MDR MRSA strain that carries the SCCmec type IV.
Sheba Medical Center is a 1,500-bed, tertiary care hospital in which ≈10,000 infants are delivered annually. MDR MRSA is endemic in the hospital, constituting 50%–60% of all S. aureus isolates, while non-MDR MRSA has been observed in only 2 cases in the last 5 years. The premature neonatal department admits 500 premature neonates annually, nearly 100% of whom are born in the hospital. The department contains ≈45 beds: 12–14 level 3 NICU beds in a single large space, 15 intermediate-intensive beds in a separate room, and 18 additional beds in 2 additional rooms. The NICU is separated by a corridor from the rest of the department. The nurse/patient ratio in the NICU is 1:3–1:5.
During October 2003, the microbiology laboratory identified MRSA blood isolates from 2 neonates in the NICU that were atypically susceptible to erythromycin, clindamycin, gentamicin, rifampicin (rifampin), trimethoprim-sulfamethoxazole (TMP-SMX), and ofloxacin (therefore designated non-MDR MRSA). Because of the rarity of such a phenotype in this institution, a retrospective search for all S. aureus clinical isolates from the premature neonatal department since 1998 was undertaken by using computerized laboratory data.
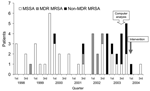
A total of 14 neonatal infections with non-MDR MRSA were discovered during this period. A cluster of 12 cases was observed from October 2002 to December 2003 (designated the outbreak period), while 2 sporadic cases were isolated from January 1, 1998, to September 31, 2002, the preoutbreak period (Figure 1). The incidence of all S. aureus infections was 15.2 per 1,000 hospitalized neonates in the preoutbreak period, while during the outbreak period, it increased to 27.7 (p = 0.032). The incidence of non-MDR MRSA was 1.4 cases per 1,000 hospitalized neonates in the preoutbreak period; the rate increased to 18.5 cases per 1,000 hospitalized neonates (p < 0.0001) during the outbreak period (Table).
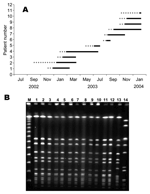
The first case of the cluster occurred 15 months after the second sporadic case. This case was in a 690-g female neonate of 25 weeks’ gestational age, hospitalized in the NICU since birth. A non-MDR MRSA was isolated from eye discharge on her 22nd day of life, and she recovered with no antimicrobial drug treatment. The next 11 case-patients (Figure 2A) were also hospitalized in the NICU since birth; 9 were bacteremic and had signs of sepsis, 4 had sputum isolates (2 of these patients had pneumonia), 2 died, and 1 death was attributable to the infection. Thirteen of 14 neonates were treated with vancomycin for periods ranging from 6 to 48 days; 2 were treated with vancomycin and rifampicin. None of the patients had perinatal infections, and the median age when infection was diagnosed was 30 days (range 11–115 days). Eleven patients had extremely low birth weight (range 568–2,440 g, median 810 g), and 12 had low gestational age (range 23–35 weeks, median 25 weeks). All patients had indwelling devices (mechanical ventilation for 11 to 74 days, central lines and total parenteral nutrition for 7–38 days). Four neonates had previous major surgery, and all neonates had at least 1 of several complications of prematurity (respiratory distress syndrome, bronchopulmonary dysplasia, intraventricular hemorrhage, retinopathy of prematurity).
An infection control intervention was initiated on December 5, 2003. Screening for S. aureus carriage was performed by nasal and umbilical sampling of all hospitalized neonates (N = 30; all were sampled on day 1) and nasal sampling of all the department healthcare workers (N = 114 [47 nurses, 30 physicians, 37 other healthcare workers]; 85% were sampled within 5 days). Swabs were streaked onto a differential media (CHROMagar plates, HyLabs, Rehovot, Israel) and incubated for 24 to 48 hours at 35°C. Suspicious colonies were then conventionally identified. Oxacillin resistance was determined according to National Committee for Clinical Laboratory Standards recommendations (6) and was verified by polymerase chain reaction (PCR) for the presence of mecA (1). Eight patient-unique, non-MDR S. aureus blood isolates from the previous year were also available for analyses.
In this point-prevalence surveillance, a non-MDR MRSA was isolated from 3 neonates with previous clinical isolates and from 4 newly recognized carriers. Thus, 7 (23.3%) of 30 hospitalized neonates were carriers. In addition, 2 (1.7%) healthcare workers, both nurses (4.3% of nurses), carried this strain; MDR MRSA was not isolated. Methicillin-susceptible S. aureus was carried by 28.1% of healthcare workers and was not isolated from any of the neonates.
Pulsed-field gel electrophoresis (PFGE) was performed, as previously described (7), on 13 unique isolates from 11 neonates and 2 nurses. A single identical outbreak strain was identified by the PFGE pattern (Figure 2B). All non-MDR MRSA were found to be SCCmec type IV by a modification of the method described by Oliveira (8). By using multilocus sequence typing (MLST) (9), the outbreak strain was classified as ST45-MRSA-IV. PCR screening of the outbreak strain for 25 virulence factor genes was performed as described by Jarraud et al. (10). MLST and PCR of virulence factors were performed on 8 outbreak isolates. Virulence factors commonly found in CA-MRSA, lukS-PV-lukF-PV encoding PVL components S and F, γ-hemolysin variant (hlg-v), and agr type 3, were not detected in the outbreak strain. The outbreak strain was positive for enterotoxin gene cluster (egc), γ-hemolysin (hlg), and agr type 1 and had similar PFGE and MLST results as the major methicillin-susceptible S. aureus (MSSA) strain found in the community served by this hospital. The outbreak strain was identical to the CA-MRSA that is rarely isolated in healthy carriers in Israel (11). Moreover, this strain differed from the MDR MRSA strains that are commonly isolated in this medical center (see Figure 2B).
After initial surveillance, all case-patients were isolated and cohorted, appropriate hand hygiene practices were reinforced for healthcare workers, and all neonates were bathed with diluted (1:10) chlorhexidine gluconate 4% once daily for 3 consecutive days. Nasal mupirocin was implemented 3 times per day for 5 consecutive days for all carriers. These regimens were well tolerated with no adverse events.
Three weeks after the first surveillance and intervention, a second surveillance was conducted in the NICU and intermediate neonatal care unit. No new cases were identified. Of the 7 carriers, only 4 were still hospitalized. Of these, 2 continued to carry the outbreak strain, and 2 were successfully decolonized. The 2 carriers were decolonized after a second course of a similar regimen (chlorhexidine bath followed by mupirocin administration).
The 2 colonized nurses were sampled twice, 1 week apart, and were found to be persistent carriers of the outbreak strain. They were instructed about good hand hygiene practices, and nasal mupirocin was recommended for 5 days. One nurse cleared her nasal non-MDR MRSA after mupirocin treatment and acquired a new strain of MSSA 2 weeks later. The other refused mupirocin treatment and persistently carried the outbreak strain for 8 weeks. During this period, despite our instructions, she continued to work until she went on maternity leave. Before she returned to work 12 weeks later, she was decolonized by using nasal mupirocin. In a 7-month follow-up, no new cases of non-MDR MRSA were identified (Figure 1).
This report documents a nosocomial outbreak of a non-MDR, PVL-negative MRSA strain, ST45-IV, in a NICU. This strain is clearly distinct from the NA-MRSA strains in this medical center, but it is identical to a CA-MRSA strain previously isolated in the community (in 0.5% of S. aureus carriers) and is nearly identical to the major MSSA strain circulating in this community (11). We hypothesize that the outbreak strain evolved in the community and penetrated into the NICU. Two previous reports of possible CA-MRSA in NICUs have been reported; however, neither characterized the alleged pathogen genetically (12,13).
The outbreak strain reported here is similar to CA-MRSA strains described in Europe, the United States, and Australia in that it is susceptible to many antimicrobial drugs and carries the SCCmec type IV (2,5). However, this strain is distinct from those strains because it lacks the PVL components S and F, as well as other virulence factors. Indeed, skin and soft-tissue infections typically described as caused by CA-MRSA (1,2) were not observed in any of the neonates in this outbreak.
MRSA outbreaks in NICUs have been reported to be difficult to contain (12,14). Only implementation of aggressive infection control measures, frequently combined with mupirocin treatment, has been successful in controlling such outbreaks. The outbreak described here was similarly contained by implementing a multifaceted infection control intervention. Since all the measures were undertaken simultaneously, we cannot define which of the measures was the most important.
We assume that the primary source of this outbreak was either a parent of the index patient or a carrier nurse. Because of a low ratio of nurses to neonates in the NICU and the high contact rate each nurse had with neonates in that facility, we could not trace specific contact patterns or perform a case-control study to further investigate this outbreak.
The difficulties encountered in implementing proper infection control measures are demonstrated by the fact that 1 healthcare worker continued to be engaged in patient care for 8 weeks, despite continued colonization by the outbreak strain, against our advice to her supervisors. Fortunately, this action did not result in persistence of the outbreak.
This outbreak illustrates the penetration of a community pathogen into the hospital, where nosocomial transmission, particularly in an intensive care setting, may rapidly spread the pathogen. The susceptibility of newborns, coupled with insufficient infection control measures and inadequate nurse-to-patient ratio, contributed to this outbreak. We call for specific attention to the possibility of “reverse penetration” of community MRSA strains becoming nosocomial pathogens.
Dr. Regev-Yochay is currently a research fellow at the Harvard School of Public Health, Boston, Massachusetts. She conducts research in the field of bacterial interference; antimicrobial resistance in the community, including transmission of resistant bacteria in families; judicious microbial treatment; and prevalence of resistant Streptococcus pneumoniae, Staphylococcus aureus, and Escherichia coli in the community.
Acknowledgment
We thank the microbiology laboratory staff for the intense effort they made and the premature neonatal ward staff for their full cooperation.
References
- Herold BC, Immergluck LC, Maranan MC, Lauderdale DS, Gaskin RE, Boyle-Vavra S, Community-acquired methicillin-resistant Staphylococcus aureus in children with no identified predisposing risk. JAMA. 1998;279:593–8. DOIPubMedGoogle Scholar
- Naimi TS, LeDell KH, Como-Sabetti K, Borchardt SM, Boxrud DJ, Etienne J, Comparison of community- and health care-associated methicillin-resistant Staphylococcus aureus infection. JAMA. 2003;290:2976–84. DOIPubMedGoogle Scholar
- Salgado CD, Farr BM, Calfee DP. Community-acquired methicillin-resistant Staphylococcus aureus: a meta-analysis of prevalence and risk factors. Clin Infect Dis. 2003;36:131–9. DOIPubMedGoogle Scholar
- Daum RS, Ito T, Hiramatsu K, Hussain F, Mongkolrattanothai K, Jamjlang M, A novel methicillin-resistance cassette in community-acquired MRSA isolates of diverse genetic backgrounds. J Infect Dis. 2002;186:1344–7. DOIPubMedGoogle Scholar
- Vandenesch F, Naim T, Enright MC, Lina G, Nimo GR, Heffernan H, Community-acquired methicillin-resistant Staphylococcus aureus carrying Panton-Valentine leukocidin genes: worldwide emergence. Emerg Infect Dis. 2003;9:978–84.PubMedGoogle Scholar
- National Committee for Clinical Laboratory Standards. Performance standards for antimicrobial susceptibility testing. NCCLS M100-S14. 2004.
- Tenover F, Arveit RD, Goering RV, Pickelsen PA, Murray BE, Persing DH, Interpreting chromosomal DNA restriction patterns produced by pulsed-field gel electrophoresis: criteria for bacterial strain typing. J Clin Microbiol. 1995;33:2233–9.PubMedGoogle Scholar
- Oliveira DC, de Lencastre H. Multiplex PCR strategy for rapid identification of structural types and variants of the mec element in methicillin-resistant Staphylococcus aureus. Antimicrob Agents Chemother. 2002;46:2155–61. DOIPubMedGoogle Scholar
- Enright MC, Robinson DA, Randle G, Feil EJ, Grundmann H, Spratt BG. The evolutionary history of methicillin-resistant Staphylococcus aureus in a UK hospital. Proc Natl Acad Sci U S A. 2002;99:7687–92. DOIPubMedGoogle Scholar
- Jarraud S, Mougel C, Thioulouse J, Lina G, Meugnier H, Forey F, Relationships between Staphylococcus aureus genetic background, virulence factors, agr groups (alleles) and human disease. Infect Immun. 2002;70:631–41. DOIPubMedGoogle Scholar
- Regev-Yochay G, Carmeli Y, Raz M, Shainberg B, Derazne E, Schwaber M, Prevalence and genetic relatedness of community MRSA strains in central Israel [abstract #1975]. In: Abstracts of the 43rd Interscience Conference on Antimicrobial Agents and Chemotherapy; 2003 Sep 14–17; Chicago, IL. Washington: American Society for Microbiology; 2003.
- Andersen BM, Lindemann R, Bergh K, Neheim BI, Syversen G, Solheim N, Spread of methicillin-resistant Staphylococcus aureus in a neonatal intensive unit associated with understaffing, overcrowding and mixing of patients. J Hosp Infect. 2002;50:18–24. DOIPubMedGoogle Scholar
- Morel AS, Wu F, Della-Latta P, Cronquist A, Rubenstein D, Saiman L. Nosocomial transmission of methicillin-resistant Staphylococcus aureus from a mother to her preterm quadruplet infants. Am J Infect Control. 2002;30:170–3. DOIPubMedGoogle Scholar
- Saiman L, Cronquist A, Wu F, Zhou J, Rubenstein D, Eisner W, An outbreak of methicillin-resistant Staphylococcus aureus in a neonatal intensive care unit. Infect Control Hosp Epidemiol. 2003;24:317–21. DOIPubMedGoogle Scholar
Figures
Table
Cite This ArticleTable of Contents – Volume 11, Number 3—March 2005
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Gili Regev-Yochay, Infectious Disease Unit, Sheba Medical Center, Tel-Hashomer, Ramat-Gan, 52621 Israel; fax: 972-3-5303501
Top