Volume 11, Number 7—July 2005
Dispatch
Caliciviruses and Foodborne Gastroenteritis, Chile
Figure
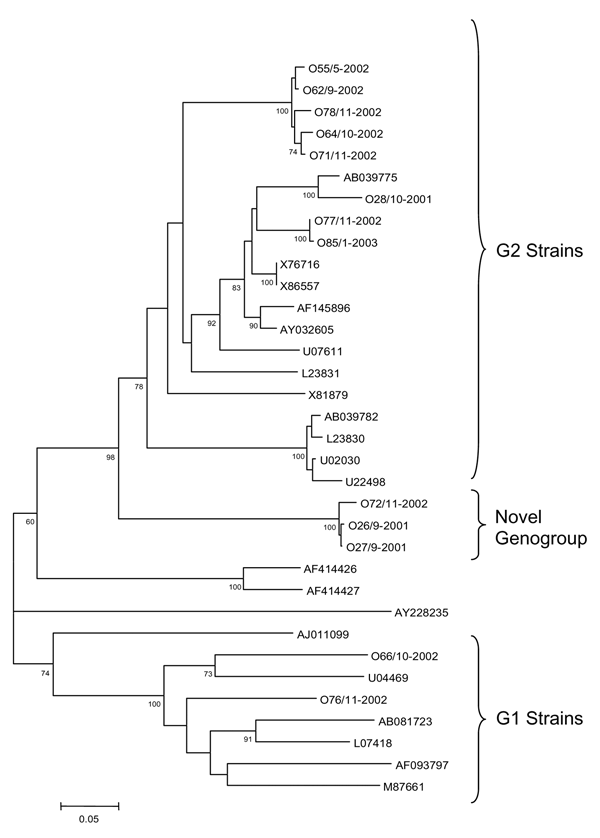
Figure. . Phylogenetic tree of noroviruses based on the 327-base region of the 3´ end of the open reading frame 1 using 13 novel sequences designated according to outbreak number/month–year (example: O55/5–2002), and 21 sequences of Norwalk-like virus strains representative of the currently identified genogroups, designated according to GenBank accession number. Comparative strains include: Norwalk virus (M87661), SaitamaU1 (AB039775), Saitama U201 (AB039782), WUG1 (AB081723), Schreier (AF093797), Camberwell (AF145896), Fort Lauderdale (AF414426), Saint Cloud (AF414427), Jena (AJ011099), Maryland (AY032605), Murine NV (AY228235), Southampton (L07418), OTH25 (L23830), Snow Mountain (L23831), Toronto (U02030), Desert Shield virus (U04469), Hawaii (U07611), Mexico (U22498), Bristol (X76716), Melksham (X81879), and Lorsdale (X86557). Bootstrap values based on 1,000 generated trees are displayed at the nodes (values >60% are shown).