Volume 12, Number 12—December 2006
Research
Transmission of Human and Macaque Plasmodium spp. to Ex-Captive Orangutans in Kalimantan, Indonesia
Figure 2
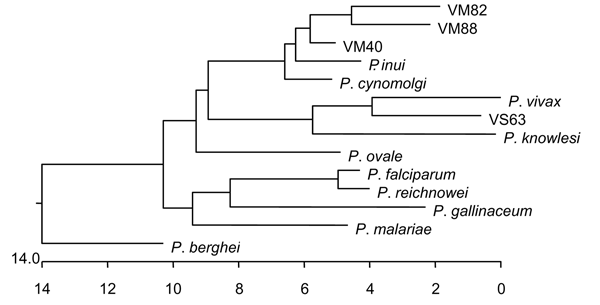
Figure 2. Phylogenetic tree of small subunit ribosomal RNA from different Plasmodium spp. Sequences were downloaded from GenBank, aligned by using CLUSTAL W (Megalign, DNA Star, Madison, WI, USA) and the tree generated by nearest-neighbor analysis. Once the sequences were aligned, we also aligned our representative sequences with the 2 nearest matches for more detailed determination of closest associations. Sequences used and their GenBank accession nos. were P. gallinaceum (M61723), P. berghei (AJ243513), P. falciparum (AL929354), P. ovale (AJ001527), P. malariae (AF88000), P. vivax (U03080), P. cynomolgi (L08241), P. fragile (M61722), P. knowlesi (U83876), P. reichenowi (Z25819), P. simium (U69605), and P. inui (U72541), clone 40 (P. cynomolgi [DQ660816]), clone 63 (P. vivax [DQ660817]), clone 82 (P. cynomolgi–like [DQ660818]), and clone 88 (P. inui–like [DQ660819]).
1Current affiliation: University of Toronto, Toronto, Ontario, Canada.
2Current affiliation: Tacugama Chimpanzee Sanctuary, Freetown, Sierra Leone