Volume 12, Number 6—June 2006
Dispatch
Hantaviruses in Serbia and Montenegro
Figure 2
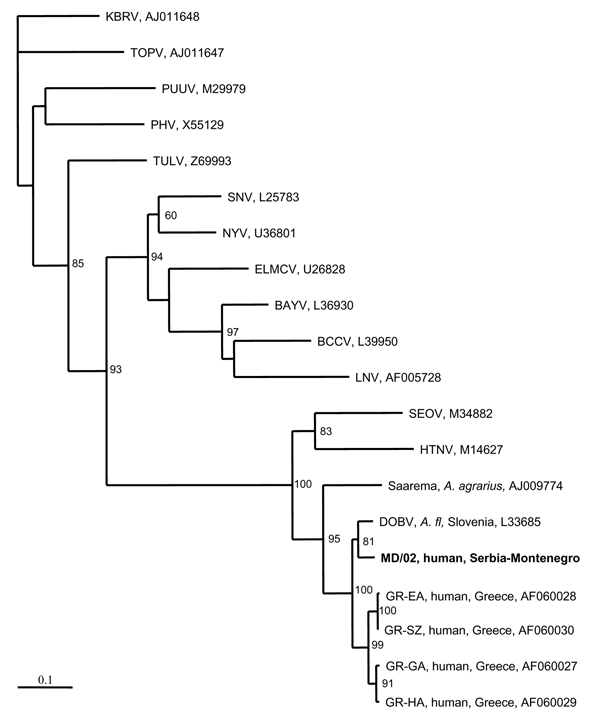
Figure 2. Phylogenetic tree based on partial M segment fragment showing the clustering of the sequence obtained from this study and respective representative hantavirus strains from GenBank database. The numbers indicate percentage bootstrap replicates (of 100); values <60% are not shown. Horizontal distances are proportional to the nucleotide differences. The scale bar indicates 10% nucleotide sequence divergence. Vertical distances are for clarity only. Sequences in this study are indicated in boldface. BAYV, Bayou virus; BCCV, Black Creek Canal virus; ELMCV, El Moro Canyon virus; HTNV, Hantaan virus; KBRV, Khabarovsk virus; LNV, Laguna Negra; MONV, Monongahela virus; NYV, New York virus; PHV, Prospect Hill virus; PUUV, Puumala virus; RIOSV, Rio Segundo virus; SEOV, Seoul virus; SNV, Sin Nombre virus; TOPV, Topografov virus; TULV, Tula virus; VLDV, Vladivostok virus.