Volume 13, Number 11—November 2007
Dispatch
Newfound Hantavirus in Chinese Mole Shrew, Vietnam
Abstract
Sequence analysis of the full-length medium segment and the partial small and large segments of a hantavirus, detected by reverse transcription–PCR in lung tissues of the Chinese mole shrew (Anourosorex squamipes) captured in Cao Bang Province, Vietnam, in December 2006, indicated that it is genetically distinct from rodentborne hantaviruses.
Insectivores (or soricomorphs) have been largely ignored as being important in the evolutionary dynamics of hantaviruses, despite the isolation of Thottapalayam virus (TPMV) from the Asian house shrew (Suncus murinus) (1,2) and the detection of hantavirus antigens in tissues of the Eurasian common shrew (Sorex araneus), alpine shrew (S. alpinus), Eurasian water shrew (Neomys fodiens), and common mole (Talpa europea) (3). Recently, genetically distinct hantavirus sequences have been found by reverse transcription–PCR in the Therese shrew (Crocidura theresae) in Guinea (4) and the northern short-tailed shrew (Blarina brevicauda) in the United States (5). In addition, a phylogenetically distinct hantavirus has been isolated from lung tissues of the Ussuri shrew (C. lasiura), captured along the Imjin River near the demilitarized zone in South Korea (J.-W. Song and R. Yanagihara, unpub. data).
To further investigate the existence and phylogeny of nonrodentborne hantaviruses, we analyzed lung and other visceral tissues, collected in RNAlater Stabilization Reagent (QIAGEN, Valencia, CA, USA), from 24 soricomorphs, including 9 white-toothed shrews (Crocidura spp.), 3 Chinese mole shrews (Anourosorex squamipes), and 12 long-nosed moles (Euroscaptor longirostris), captured in northern, central, and southern Vietnam during November and December 2006. RNA, extracted from 20–50 mg of each tissue by using the RNA-Bee isolation kit (TEL-TEST, Inc., Friendswood, TX, USA), was reverse transcribed by using Moloney murine leukemia virus reverse transcriptase (Promega, Madison, WI, USA) and the primer 5′-TAGTAGTAGACTCC-3′. Oligonucleotide primers for subsequent nested PCR were designed from consensus regions of TPMV and other hantaviruses (Table 1).
Gene-amplification reactions were performed in 50-μL reaction mixtures, containing 200 μmol deoxyribonucleoside triphosphate, 0.5 U of super-therm polymerase (PureTech Co., Ltd, Seoul, South Korea), 1 μg of cDNA, and 10 pmol of each primer. Initial denaturation, at 94°C for 5 min, was followed by touchdown cycling with denaturation at 94°C for 40 s, annealing from 50°C to 37°C for 40 s, elongation at 68°C for 1 min 20 s, then 25 cycles of denaturation at 94°C for 40 s, annealing at 40°C for 40 s, and elongation at 68°C for 1 min 20 s in a Mastercycler ep gradient S (Eppendorf AG, Hamburg, Germany). PCR products were purified by the Wizard PCR Preps DNA Purification System (Promega). DNA sequencing of at least 3 clones of each amplicon was performed in both directions with the dye primer cycle sequencing ready reaction kit (Applied Biosystems, Foster City, CA, USA) on an automated sequencer (Model 377, Perkin Elmer Co., Waltham, MA, USA) (6).
Hantavirus sequences were not detected in tissues of the white-toothed shrews and long-nosed moles. By contrast, the full-length 3,637-nt (1,139-aa) medium (M) segment was amplified from lung tissues of 3 Chinese mole shrews, captured in Thanh Cong commune, Nguyen Binh District, Cao Bang Province, along the southern border of the People’s Republic of China. Designated Cao Bang virus (CBNV), the newly identified hantavirus exhibited low nucleotide and amino acid sequence similarity to representative hantaviruses harbored by Murinae, Arvicolinae, Neotominae, and Sigmodontinae rodents, ranging from 62.6% (nt) and 61.2% (aa) for Hantaan virus (HTNV) 76–118; 62.3% (nt) and 61.7% (aa) for Dobrava virus (DOBV) Greece to 58.1% (nt) and 52.0% (aa) for Puumala virus Sotkamo; and 58.8% (nt) and 54.7% (aa) for Sin Nombre virus NMH10 (Table 2).
Pairwise alignment and comparison of a 1,185-nt coding region of the small (S) segment showed similar degrees of sequence identity between CBNV and rodentborne hantaviruses, ranging from 61.4% for HTNV 76–118 to 58.0% for TULV (Tula virus) M5302v. Much higher sequence similarity was found in a 412-nt coding region of the large (L) segment, ranging from 72.6% for HTNV 76–118 and 75.2% for DOBV Greece, to 67.2%–70.1% and 67.7%–71.6% for hantaviruses harbored by Arvicolinae, Neotominae, and Sigmodontinae rodents, respectively. CBNV sequences were similarly divergent from Tanganya virus (TGNV) Tan826: for S segment, 63.5% (nt) and 65.0% (aa) similarity; for L segment, 71.3% (nt) and 77.0% (aa) similarity.
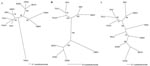
Phylogenetic trees based on sequences of the full-length M segment and partial S and L segments, generated by the maximum likelihood and neighbor-joining methods using the GTR+I+G model of evolution, showed similar topologies supported by bootstrap analysis, in which CBNV was relatively distinct from rodentborne and other shrewborne hantaviruses (Figure). A strong association with TGNV was observed on the basis of the S segment (1,185 bases), however. Further sequence information will clarify the relationship between CBNV and other soricidborne hantaviruses and whether these form a monophyletic group in parallel with the evolution of Soricinae and Crocidurinae shrews. If one judges by the distant evolutionary relationship between shrews and rodents, future sequences of other soricidborne hantaviruses will provide considerable insights into their evolutionary origins.
Designing suitable primers for the amplification of CBNV presented unanticipated challenges. Ironically, the recently acquired full genome of TPMV (J.-W. Song and R. Yanagihara, unpub. data) was not particularly helpful, since CBNV was genetically more divergent from TPMV than from well-characterized rodentborne hantaviruses (78% bootstrap support, S segment). Also, because the tissues were collected in RNAlater, virus isolation attempts could not be performed. As such, progress in obtaining the full-length sequence of CBNV has been slow.
A forest-dwelling soricine typically residing at elevations of 1,500–3,000 m, the Chinese mole shrew (family Soricidae, subfamily Soricinae) has a vast geographic range, extending from western and central People’s Republic of China, northern Myanmar, northern Thailand, Assam, Bhutan, northern Vietnam, Taiwan, and possibly Lao People’s Democratic Republic. That a hantavirus has been identified in the Chinese mole shrew was not completely unexpected, in view of the isolation of a HTNV-like virus from this species in Sichuan Province in 1986 (7). However, those authors may have prematurely concluded that their hantavirus isolate was closely related to HTNV, since no genetic analysis was performed.
Viewed within the context of newly identified, genetically distinct hantaviruses in the northern short-tailed shrew (B. brevicauda), Eliot’s short-tailed shrew (B. hylophaga), masked shrew (Sorex cinereus), and montane shrew (S. monticolus) in the United States (S. Arai and R. Yanagihara, unpub. data)—as well as in the Eurasian common shrew in Switzerland, Hungary, and Finland (J.-W. Song, S. Arai, and R. Yanagihara, unpub. data), the Ussuri shrew in South Korea (J.-W. Song and R. Yanagihara, unpub. data), the Asian house shrew in India (1,2), and the Therese shrew in Guinea (4)—the detection of a newfound hantavirus in the Chinese mole shrew would predict that hantaviruses harbored by shrews are as geographically widespread as those harbored by rodents. Preliminary studies indicate CBNV-like sequences in the liver tissue of Chinese mole shrews captured in Taiwan (S. Arai and R. Yanagihara, unpub. data). Also, investigations on the genetic diversity of CBNV and other newly identified members of the Hantavirus genus will provide additional insights into the phylogeography and co-evolution of hantaviruses and their soricid reservoir hosts. One or more of these newfound shrewborne viruses may yield valuable clues about the molecular determinants of hantavirus pathogenesis.
Dr Song is a professor of microbiology at Korea University. He has had a long-standing research interest in the discovery and characterization of new hantaviruses.
Acknowledgment
This work was supported in part by grants P20RR018727 (Centers of Biomedical Research Excellence) and G12RR003061 (Research Centers in Minority Institutions) from the National Center for Research Resources, National Institutes of Health, and by grant R21-2005-000-10017-0 from MOST (KOSEF), South Korea.
References
- Carey DE, Reuben R, Panicker KN, Shope RE, Myers RM. Thottapalayam virus: a presumptive arbovirus isolated from a shrew in India. Indian J Med Res. 1971;59:1758–60.PubMedGoogle Scholar
- Song J-W, Baek LJ, Schmaljohn CS, Yanagihara R. Thottapalayam virus, a prototype shrewborne hantavirus. Emerg Infect Dis. 2007;13:980–5.PubMedGoogle Scholar
- Yanagihara R, Gajdusek DC. Hemorrhagic fever with renal syndrome: a historical perspective and review of recent advances. In: Gear JHS, editor. CRC handbook of viral and rickettsial hemorrhagic fevers. Boca Raton (FL): CRC Press, Inc.; 1988. p. 151–88.
- Klempa B, Fichet-Calvet E, Lecompte E, Auste B, Aniskin V, Meisel H, Novel hantavirus sequences in shrew, Guinea. Emerg Infect Dis. 2007;13:520–2. DOIPubMedGoogle Scholar
- Arai S, Song J-W, Sumibcay L, Bennett SN, Nerurkar VR, Parmenter C, Hantavirus in northern short-tailed shrew, United States. Emerg Infect Dis. 2007;13:1420–3.PubMedGoogle Scholar
- Baek LJ, Kariwa H, Lokugamage K, Yoshimatsu K, Arikawa J, Takashima I, Soochong virus: a genetically distinct hantavirus isolated from Apodemus peninsulae in Korea. J Med Virol. 2006;78:290–7. DOIPubMedGoogle Scholar
- Chen SZ, Chen LL, Tao GF, Fu JL, Zhang CA, Wu YT, Strains of epidemic hemorrhagic fever virus isolated from the lungs of C. russula and A. squamipes [in Chinese]. Zhonghua Yu Fang Yi Xue Za Zhi. 1986;20:261–3.PubMedGoogle Scholar
Figure
Tables
Cite This ArticleTable of Contents – Volume 13, Number 11—November 2007
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Richard Yanagihara, Pacific Center for Emerging Infectious Diseases Research, John A. Burns School of Medicine, University of Hawaii at Manoa, 651 Ilalo St, BSB320L, Honolulu, HI 96813, USA;
Top