Volume 13, Number 12—December 2007
Research
Emergence of Methicillin-Resistant Staphylococcus aureus of Animal Origin in Humans
Cite This Article
Citation for Media
Abstract
In 2003 in the Netherlands, a new methicillin-resistant Staphylococcus aureus (MRSA) strain emerged that could not be typed with Sma1 pulsed-field gel electrophoresis (NT-MRSA). The association of NT-MRSA in humans with a reservoir in animals was investigated. The frequency of NT-MRSA increased from 0% in 2002 to >21% after intensified surveillance was implemented in July 2006. Geographically, NT-MRSA clustered with pig farming. A case–control study showed that carriers of NT-MRSA were more often pig or cattle farmers (pig farmers odds ratio [OR] 12.2, 95% confidence interval [CI] 3.1–48.6; cattle farmers OR 19.7, 95% CI 2.3–169.5). Molecular typing showed that the NT-MRSA strains belonged to a new clonal complex, ST 398. This study shows that MRSA from an animal reservoir has recently entered the human population and is now responsible for >20% of all MRSA in the Netherlands.
Methicillin-resistant Staphylococcus aureus (MRSA) has traditionally been considered a nosocomial pathogen. However, for several years the number of reports of so-called community-onset MRSA (CO-MRSA) has been rapidly increasing (1). CO-MRSA has no relation to healthcare and is usually associated with the presence of Panton-Valentine leukocidin toxin (PVL) and SCCmec types IV and V (2,3). In 2004 and 2005, some unexpected cases of MRSA were found in patients who were associated with pig farms (4,5). Genotyping showed that these MRSA isolates were nontypable by pulsed-field gel electrophoresis (PFGE) and belonged to 1 spa type (t108). The aims of this study were to determine if nontypable MRSA (NT-MRSA) isolates are associated with pig farming and to compare the phenotypic, genotypic, and epidemiologic features of NT-MRSA with those of typable MRSA strains.
National MRSA Database
The National Institute for Public Health and the Environment (RIVM) is the national reference center for MRSA in the Netherlands (www.rivm.nl/mrsa). According to national guidelines, all microbiology laboratories send the first isolate of newly identified carriers of MRSA to RIVM. Strains are confirmed to be MRSA by a Martineau PCR and by mecA PCR assay (6,7). Since 2002, all strains are typed by using PFGE (8), and the presence of PVL genes is determined (9).
Selection of Cases and Controls
Cases and controls were selected from the national MRSA database at RIVM. Case-patients were defined as persons carrying NT-MRSA who provided the first isolate from a cluster of 1 particular referring laboratory (index-patient) in the period January 2003 to September 2005. Cases were considered to be secondary to an index case when the strain was isolated within 3 months after the previous isolate with the same PFGE typing result. Controls were persons who carried MRSA that was typable with PFGE and who also fulfilled the index-patient definition. Controls were derived from the laboratories that provided cases and were selected at random. Twice as many controls as case-patients were selected.
Collection of Epidemiologic Background Information
Data were collected by questionnaires that were sent to the referring laboratories. The questionnaire contained items about patient characteristics (birth date, sex, postal code, presence or absence of infection, hospital admission dates, profession, profession of partner, profession of parents, and contact with animals, e.g., pigs, cows, horses, chickens, cats, or dogs) and microbiologic data (isolation date, source of culture, medical specialty). All data were collected and entered into the database without our knowing whether it concerned a case or control.
Initially, 41 cases and 82 controls from 26 different laboratories were selected from the national database. The response rate was 98% (40 cases and 81 controls). During workup, 5 cases and 5 controls were excluded for the following reasons: the confirmation test of the isolate indicated that it was not methicillin resistant (1 case), or the case did not fulfill the case definition because it was not the first case from a cluster (4 cases). Since 2 of these cases were from laboratories that had no other case in the study, the accompanying controls were excluded (n = 3). Two controls were identified outside the study period. Finally, 35 cases and 76 controls from 24 different laboratories were included in the analysis.
Molecular Typing and Susceptibility Testing
All MRSA isolates were typed by PFGE (8). All isolates from case-patients and 74 isolates from controls were typed by spa typing (10). Multilocus sequence typing (MLST) was performed on all case isolates, as well as on 1 strain of each spa type of the control isolates (n = 37) (11). PCR of the staphylococcal chromosome cassette (SCCmec) was performed according to Zhang et al. on all isolates from case-patients and 74 isolates from controls (12). The presence or absence of PVL genes (lukS-PV/lukF-PV) was determined in all case isolates and in 71 control isolates. The PVL genes were detected by PCR according to the method of Lina et al. (13). The susceptibility to antimicrobial agents was tested for 32 case isolates and 74 control isolates, according to CLSI guidelines that used Mueller-Hinton agar and multipoint inoculation (14).
Statistical Analysis
Data were entered into an Excel database (Microsoft Windows version 97 SR-2, Redmond, WA, USA) and further analyzed by using SAS (version 9.1) software package (SAS Institute Inc., Cary, NC, USA). Chi-square test and Fisher exact test for ordinal variables and Student t test for continuous variables were used for univariate analysis. Variables associated with both case-control status and the exposure (i.e., contact with pigs or cattle, respectively) with a p value <0.2 were included in the multivariate logistic regression model. If such variables changed the risk estimate for >10%, they were left in the model. All statistical tests were 2-sided, and a p value <0.05 was considered statistically significant.
Epidemic Curve
The first NT-MRSA isolate was found in February 2003. In subsequent years, an increasing number of NT-MRSA isolates were found. The percentage of NT-MRSA relative to the total number of MRSA isolates in the Netherlands that were unique or first from a cluster rose from 0% in 2002 to 5.5% in the first half of 2006 and to >21% in the second half of 2006, after the introduction of intensified surveillance in July 2006.
Geographic Distribution
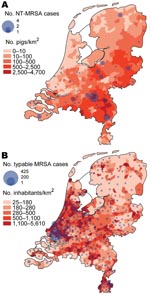
Figure 1 shows the geographic distribution of NT-MRSA and typable MRSA isolates, plotted over the density of the pig and human populations, respectively. The density of NT-MRSA isolates corresponds to the density of pig farming, whereas the density of typable strains corresponds to the density of the human population. The density of cattle farms is more or less identical to the density of pig farms.
Epidemiologic Data
Results of the univariate analysis are shown in Table 1. Comparable values were observed for the baseline characteristics of sex and age. Case-patients more often lived in rural areas and indicated more frequent contact with pigs or cattle than did controls. Controls were more often associated with healthcare facilities.
Among case-patients, MRSA was more frequently found in clinical samples (an unexpected finding) compared with controls, whose MRSA was found more often by targeted screening in nose, throat, and perineum. Among persons infected by MRSA, respiratory tract infections were more frequent in case-patients, whereas skin and soft tissue infections predominated in controls.
Multivariate analysis that used a model with the variables describing type of residence (rural vs. other) and contact with pigs, cattle, cats, and dogs (yes, no, or unknown) showed that contact with pigs and contact with cattle were independent statistically significant variables. The adjusted odds ratios (OR) for pigs and cattle were 9.4 (95% confidence interval [CI] 1.8–47.7) and 13.5 (95% CI 1.0–179.3), respectively.
Molecular Typing
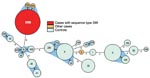
Thirty-two of 35 case-patients had MLST sequence type (ST) 398; 1 had ST 9; and the remaining 2 had ST 752 and 753, closely related to 398 (Figure 2). Among case-patients, the most frequent spa types were t108, t011, and t034 (Table 2). These MLST and spa types were not found among the controls. Twenty-two different STs and 37 different spa types were found in the controls (Table 2 and Figure 2).
SCCmec typing showed that in isolates from cases SCCmec types III, IV, and V were found, whereas in isolates from controls all SCCmec types were found (Table 2). For 11 cases and 33 controls, the SCCmec type could not be determined. There was no difference in the presence of the PVL genes (Table 2).
Antimicrobial Agent Susceptibility
Table 3 shows the percentage of strains that were resistant to various antimicrobial agents. Isolates from case-patients were significantly more often resistant to doxycycline and clindamycin than were isolates from controls.
A new type of MRSA recently emerged in the Netherlands. The first isolate was found in 2003, and since then it has been found with increasing frequency. The geographic origin of NT-MRSA correlates with the density of pig populations. This association was confirmed by the results from this case-control study, which show that NT-MRSA is significantly related to contact with pigs. In addition, a significant association was found with cattle. After multivariate analysis, contact with pigs and cattle were the only 2 significant independent variables. Screening of a representative sample of pigs in the Netherlands was recently performed and showed that nearly 40% of the pigs were colonized with a comparable strain of MRSA (MLST 398) and that ≈80% of the pig farms were affected (15). The association between NT-MRSA and cattle was not expected when this study was initiated and needs further evaluation.
On the basis of the above-mentioned findings, we conclude that this new MRSA strain is of animal origin (pigs and probably cows). Transmission of MRSA between animals and humans has previously been described, e.g., associated with colonized companion animals, horses, and persons who take care of them (16–19). However, the MRSA clones in these reports were known human clones, suggesting human-to-animal transmission in origin. Baptiste et al. found specific PFGE clones in horses that were never observed before (20). Until now, transmission of these clones to humans has not been reported.
We assume that this problem is not limited to the Netherlands. First, widespread dissemination in pigs in the Netherlands has been found. When one considers the intensive international transport of pigs, it is unlikely that this situation is limited to the Netherlands. Second, 3 of the case-patients came from abroad, 1 tourist and 2 adopted children from Asia. Also, MLST 398 was recently found in animals (pig, dog, and foal) and in humans in Germany (21). Finally, in Hong Kong Special Administrative Region, People’s Republic of China, MRSA with MLST 398 has been found in 2 patients with bacteremia (22).
The origin of the current NT-MRSA situation is difficult to elucidate. One earlier study can be found on carriage of S. aureus in pig farmers and pigs in France (23). It reported an increased carriage rate in pig farmers caused by transmission of S. aureus from pigs that also carried MLST ST 9 and 398. Further typing of the French ST 398 isolates at RIVM showed homology with the Dutch isolates. However, in the French study most of the MLST 398 strains were susceptible to β-lactam antimicrobial agents. The most likely explanation for the current findings is that MLST 398 is a commensal strain in pigs, which originally was methicillin susceptible. As most NT-MRSA isolates were resistant to doxycycline, the spread is facilitated by the abundant use of tetracyclines in pig and cattle farming (15).
What are the implications of these findings? Persons working or living in close contact with pigs or cows are at increased risk of becoming colonized and infected with MRSA. Infections can be severe, as is indicated by the hospital admission rate. Also, a case of endocarditis has been reported recently (24). At present, whether this strain is spreading further in the community is not clear. Before final recommendations for control can be made, the current size of the reservoir in farm animals and in humans has to be determined at an international level.
Dr van Loo is a medical microbiologist at the University Hospital Maastricht. The research described in this manuscript was performed as part of her training as a medical microbiologist.
Acknowledgment
We thank Ing. H. Giesbers for constructing Figure 2; R. Bosboom, A. Demeulemeester, B. v. Dijke, B. Hendrickx, R. Hendrix, C. Hol, E. IJzerman, S. Kuijpers, A. Lampe, M.K.E. Nohlmans-Paulssen, N. Meessen, A. Möller, M. Peeters, D. Potters, P. Rietra, P. Schneeberger, F. Sebens, L. Spanjaard, A. Troelstra, C. Vandebroucke-Grauls, C. Verduin, and M. Wulf for answering the questionnaires; M. van Santen-Verheuvel and E. Spalburg for performing spa, MLST, and SCCmec typing and susceptibility testing; C. Klaassen for SCCmec typing; and L. Schouls for analysis of the MLST data.
References
- Diederen BMW, Kluytmans JAJW. The emergence of infections with community-associated methicillin resistant Staphylococcus aureus. J Infect. 2006;52:157–68. DOIPubMedGoogle Scholar
- Naimi TS, LeDell KH, Como-Sabetti K, Brochardt SM, Boxrud DJ, Etienne J, Comparison of community- and health care–associated methicillin-resistant Staphylococcus aureus infection. JAMA. 2003;290:2976–84. DOIPubMedGoogle Scholar
- Kluytmans-Vandenbergh MF, Kluytmans JA. Community-acquired methicillin-resistant Staphylococcus aureus: current perspectives. Clin Microbiol Infect. 2006;12:9–15. DOIPubMedGoogle Scholar
- Voss A, Loeffen F, Bakker J, Klaassen C, Wulf M. Methicillin-resistant Staphylococcus aureus in pig farming. Emerg Infect Dis. 2005;11:1965–6.PubMedGoogle Scholar
- Huijsdens XW, Van Dijke B, Spalburg E, van Santen-Verheuvel MG, Heck ME, Pluister GN, Community-acquired MRSA and pig-farming. Ann Clin Microbiol Antimicrob. 2006;5:26. DOIPubMedGoogle Scholar
- Martineau F, Picard FJ, Roy PH, Ouellette M, Bergeron MG. Species-specific and ubiquitous-DNA–based assays for rapid identification of Staphylococcus aureus. J Clin Microbiol. 1998;36:618–23.PubMedGoogle Scholar
- Wannet WJ, Spalburg E, Heck ME, Pluister GN, Willems RJ, de Neeling AJ. Widespread dissemination in The Netherlands of the epidemic Berlin methicillin-resistant Staphylococcus aureus clone with low-level resistance to oxacillin. J Clin Microbiol. 2004;42:3077–82. DOIPubMedGoogle Scholar
- Murchan S, Kaufmann ME, Deplano A, de Ryck R, Struelens M, Zinn CE, Harmonization of pulsed-field gel electrophoresis protocols for epidemiological typing of strains of methicillin-resistant Staphylococcus aureus: a single approach developed by consensus in 10 European laboratories and its application for tracing the spread of related strains. J Clin Microbiol. 2003;41:1574–85. DOIPubMedGoogle Scholar
- Wannet WJ, Spalburg E, Heck ME, Pluister GN, Tiemersma E, Willems RJ, Emergence of virulent methicillin-resistant Staphylococcus aureus strains carrying Panton-Valentine leucocidin genes in The Netherlands. J Clin Microbiol. 2005;43:3341–5. DOIPubMedGoogle Scholar
- Harmsen D, Claus H, Witte W, Rotganger J, Claus H, Turnwald D, Typing of methicillin-resistant Staphylococcus aureus in a university hospital setting by using novel software for spa repeat determination and database management. J Clin Microbiol. 2003;41:5442–8. DOIPubMedGoogle Scholar
- Enright MC, Day NP, Davies CE, Peacock SJ, Spratt BG. Multilocus sequence typing for characterization of methicillin-resistant and methicillin-susceptible clones of Staphylococcus aureus. J Clin Microbiol. 2000;38:1008–15.PubMedGoogle Scholar
- Zhang K, McClure JA, Elsayed S, Louie T, Conly JM. Novel multiplex PCR assay for characterization and concomitant subtyping of staphylococcal cassette chromosome mec types I to V in methicillin-resistant Staphylococcus aureus. J Clin Microbiol. 2005;43:5026–33. DOIPubMedGoogle Scholar
- Lina G, Piemont Y, Godail-Gamot F, Bes M, Peter MO, Gauduchon V, Involvement of Panton-Valentine leukocidin-producing Staphylococcus aureus in primary skin infections and pneumonia. Clin Infect Dis. 1999;29:1128–32. DOIPubMedGoogle Scholar
- National Committee for Clinical Laboratory Standards. Performance standards for antimicrobial susceptibility testing. Wayne (PA): The Committee; 2006.
- De Neeling AJ, Van den Broek MJM, Spalburg EC, Van Santen-Verheuvel MG, Dam-Deisz W, Boshuizen HC, High prevalence of methicillin resistant Staphylococcus aureus in pigs. Vet Microbiol. 2007;122:366–72. DOIPubMedGoogle Scholar
- Duquette RA, Nuttall TJ. Methicillin-resistant Staphylococcus aureus in dogs and cats: an emerging problem? J Small Anim Pract. 2004;45:591–7. DOIPubMedGoogle Scholar
- Cefai C, Ashurst S, Owens C. Human carriage of methicillin-resistant Staphylococcus aureus linked with pet dog. Lancet. 1994;344:539–40. DOIPubMedGoogle Scholar
- Loeffler A, Boag AK, Sung J, Lindsay JA, Guardabassi L, Dalsgaard A, Prevalence of methicillin-resistant Staphylococcus aureus among staff and pets in a small animal referral hospital in the UK. J Antimicrob Chemother. 2005;56:692–7. DOIPubMedGoogle Scholar
- Weese JS, Archambault M, Willey BM, Heam P, Kreiswirth BM, Said-Salim B, Methicillin-resistant Staphylococcus aureus in horses and horse personnel, 2000–2002. Emerg Infect Dis. 2005;11:430–5.PubMedGoogle Scholar
- Baptiste KE, Williams K, Willams NJ, Wattret A, Clegg PD, Dawson S, Methicillin-resistant staphylococci in companion animals. Emerg Infect Dis. 2005;11:1942–4.PubMedGoogle Scholar
- Cuney C. STrommenger B, Witte W. Emergence of MRSA of multilocus sequence type ST398 in animals and in humans. In: Program and abstracts of the 12th International Symposium on Staphylococci and Staphylococcal Infections; Maastricht, the Netherlands; 2006 Sep 3–6.
- Ip M, Yung RW, Ng TK, Luk WK, Tse C, Hung P, Contemporary methicillin-resistant Staphylococcus aureus clones in Hong Kong. J Clin Microbiol. 2005;43:5069–73. DOIPubMedGoogle Scholar
- Armand-Lefevre L, Ruimy R, Andremont A. Clonal comparison of Staphylococcus aureus isolates from healthy pig farmers, human controls, and pigs. Emerg Infect Dis. 2005;11:711–4.PubMedGoogle Scholar
- Ekkelenkamp MB, Sekkat M, Carpaij N, Troelstra A, Bonten M. Endocarditis caused by “pig-MRSA.”. Ned Tijdschr Geneeskd. 2006;150:2442–7.PubMedGoogle Scholar
Figures
Tables
Cite This ArticleTable of Contents – Volume 13, Number 12—December 2007
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Jan Kluytmans, Amphia Hospital Breda, Laboratory for Microbiology and Infection Control, PO Box 90158, 4800 RK Breda, the Netherlands;
Top