Volume 13, Number 6—June 2007
Dispatch
Bartonella Species in Blood of Immunocompetent Persons with Animal and Arthropod Contact
Cite This Article
Citation for Media
Abstract
Using PCR in conjunction with pre-enrichment culture, we detected Bartonella henselae and B. vinsonii subspecies berkhoffii in the blood of 14 immunocompetent persons who had frequent animal contact and arthropod exposure.
Attempts to isolate Bartonella sp. from immunocompetent persons with serologic, pathologic, or molecular evidence of infection are often unsuccessful; several investigators have indicated that Bartonella isolation methods need to be improved (1–4). By combining PCR and pre-enrichment culture, we detected B. henselae and B. vinsonii subspecies berkhoffii infection in the blood of immunocompetent persons who had arthropod and occupational animal exposure.
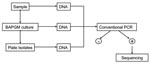
From November 2004 through June 2005, blood and serum samples from 42 persons were tested, and 14 completed a questionnaire, approved by the North Carolina State University Institutional Review Board. Age, sex, animal contact, history of bites, environment, outdoor activity, arthropod contact, travel, and medical history were surveyed. Bacterial isolation, PCR amplification, and cloning were performed by using previously described methods (5–7). Each blood sample was tested by PCR after direct DNA extraction, pre-enrichment culture for at least 7 days, and subculture onto a blood agar plate (Figure). An uninoculated, pre-enrichment culture was processed simultaneously as a control. Methods used for DNA extraction and conventional and real-time PCR targeting of the Bartonella 16S-23S intergenic spacer (ITS) region and heme-binding protein (Pap31) gene have been described (7,8). Conventional PCR amplicons were cloned with the pGEM-T Easy Vector System (Promega, Madison, WI, USA); sequencing was performed by Davis Sequencing, Inc. (Davis, CA, USA). Sequences were aligned and compared with GenBank sequences with AlignX software (Vector NTI Suite 6.0 (InforMax, Inc., Bethesda, MD, USA) (7,8). B. vinsonii subsp. berkhoffii, B. henselae, and B. quintana antibodies were determined by using a modification of a previously described immunofluorescence antibody assay (IFA) procedure (9).
Study participants included 12 women and 2 men, ranging in age from 30 to 53 years; all of them reported occupational animal contact for >10 years (Table). Most had daily contact with cats (13 persons) and dogs (12 persons). All participants reported animal bites or scratches (primarily from cats) and arthropod exposure, including fleas, ticks, biting flies, mosquitoes, lice, mites, or chiggers. All participants reported intermittent or chronic clinical symptoms, including fatigue, arthralgia, myalgia, headache, memory loss, ataxia, and paresthesia (Table). Illness was most frequently mild to moderate in severity, with a waxing and waning course, and all but 2 persons could perform occupational activities. Of the 14 participants, 9 had been evaluated by a cardiologist, 8 each by an infectious disease physician or a neurologist, and 5 each by an internist or a rheumatologist. Eleven participants had received antimicrobial drugs.
When reciprocal titers of >64 were used, 8 persons were seroreactive to Bartonella antigens (Appendix Table). B. henselae or B. vinsonii subsp. berkhoffii was detected or isolated from all 14 participants. At the time of initial testing, Bartonella DNA was amplified directly from 3 blood samples, from 7 pre-enrichment liquid cultures, and from 4 subculture isolates (Appendix Table). For 5 persons, results of PCR and culture of initial samples were negative. Overall, Bartonella DNA was amplified from 11 (28%) of 40 extracted blood samples, 13 (33%) of 40 pre-enrichment cultures, and 5 isolates. For 7 persons, B. henselae DNA was amplified at multiple time points. Bartonella DNA was never amplified from any PCR control or uninoculated culture control.
By using the ITS target region, 2 distinct B. henselae ITS and Pap31 strains were sequenced, B. henselae Houston I (HI) (GenBank NC-005956) and B. henselae San Antonio 2 (SA2) (GenBank AF369529). Within the noncoding ITS region, B. henselae SA2 strains have a 30-bp insertion (ATT GCT TCT AAA AAG ATT GCT TCT AAA AAG) located 518 bases downstream from the 16S gene. Only B. vinsonii subsp. berkhoffii types I and II were detected (8).
Persistent human infection with B. bacilliformis and B. quintana has been previously documented, whereas infection with B. henselae (cat-scratch disease [CSD]) is generally considered self-limiting (1,2,10). Recently, B. henselae DNA was amplified from the blood of a child 4 months after CSD diagnosis (11). Our study indicates that B. henselae and B. vinsonii subsp. berkhoffii can induce occult infection in immunocompetent persons and that detection can be enhanced by combining PCR with pre-enrichment culture. Considering only the results from initial blood samples, PCR detected Bartonella DNA in 3 samples, all of which were subsequently PCR positive by subculture or enrichment culture. In samples from 5 persons, pre-enrichment was necessary, and in 5 other persons, sequential sampling was necessary to detect Bartonella infection. Intermittent bacteremia, as occurs in B. henselae–infected cats (12), antimicrobial drug administration, low bacterial copy numbers, and low inoculum volume (1 mL) may have contributed to intermittent detection or inability to isolate Bartonella spp. from some participant samples. Although our approach is an improvement over historical isolation approaches, our results emphasize ongoing limitations associated with the detection of Bartonella infection. Obtaining stable Bartonella subcultures (n = 5 in this study) has proven problematic for other specialized laboratories that routinely culture for Bartonella spp. (3,4). To our knowledge, the B. vinsonii subsp. berkhoffii type II isolate described in our study is the only type II human isolate reported to date (8). Various combinations of B. henselae and B. vinsonii subsp. berkhoffii strain types were detected in the same blood sample or sequential blood samples. The coexistence of B. henselae genetic variants has been described among primary patient isolates, which suggests that multiple genotypes may emerge within the same person (13).
Overall, 57% of persons tested were seroreactive to 1 or all 3 Bartonella test antigens. Previous reports from the United States identified a B. henselae seroprevalence of 3% in healthy blood donors and a cumulative seroprevalence of 7.1% to both B. henselae and B. quintana antigens in veterinary professionals (1). In this and other studies, serologic test results did not correlate with PCR amplification or isolation results. Antigenic variability among B. henselae test strains can cause false-negative IFA results in persons with suspected CSD. Also B. henselae, B. quintana, or B. elizabethae antibodies were not detected in some persons with DNA evidence of active infection (1,3,4).
Animal contact, often to a wide spectrum of domestic and wild animal species, is an obvious consequence of the daily activities of the study population, which is biased by veterinary occupational exposure and by self-selection (volunteer bias). Cats are considered the primary reservoir host for B. henselae, whereas coyotes and foxes are considered reservoir hosts for B. vinsonii subsp. berkhoffii (1,2,8). Detection of B. vinsonii subsp. berkhoffii in 4 of 5 Californian participants could be related to the high prevalence of bacteremic coyotes in this region as well as to the potential transmission by a tick vector (1,2). All 14 participants reported frequent arthropod exposure. Although Bartonella spp.transmission by ticks has not been proven, several recent studies have identified Bartonella DNA in questing ticks, ticks attached to animals, and ticks attached to humans (1,2,14).
Despite reporting chronic or episodic illness, most participants continued to effectively maintain daily professional and personal activities. The symptoms described in the study patients are very similar to those described in a community and hospital-based surveillance study of CSD patients, in whom CSD-associated arthropathy was an uncommon chronic syndrome affecting mostly young and middle-age women (15). Our study was initiated to investigate the feasibility of combining PCR with pre-enrichment culture. Prospective studies, with appropriate controls, are needed to characterize the prevalence and clinical relevance of persistent Bartonella infection in immunocompetent persons.
Dr Breitschwerdt is a professor of medicine and infectious diseases at the College of Veterinary Medicine, North Carolina State University. He is also adjunct associate professor of medicine at Duke University Medical Center. His research focuses on comparative medical aspects of zoonotic vectorborne infections in cats, dogs, and humans.
Acknowledgments
We thank the study participants for providing blood samples, Julie Bradley and Maria Belen Cadenas for technical assistance, and Tonya Lee for editorial assistance.
This research was supported by the state of North Carolina and, in part, through a gift from Bayer Animal Health (to R.G.M. and A.W.D.).
References
- Chomel BB, Kasten RW, Sykes JE, Boulouis HJ, Breitschwerdt EB. Clinical impact of persistent Bartonella bacteremia in humans and animals. Ann N Y Acad Sci. 2003;990:267–78. DOIPubMedGoogle Scholar
- Boulouis H-J, Chang CC, Henn JB, Kasten RW, Chomel BB. Factors associated with the rapid emergence of zoonotic Bartonella infections. Vet Res. 2005;36:383–410. DOIPubMedGoogle Scholar
- La Scola B, Raoult D. Culture of Bartonella quintana and Bartonella henselae from human samples: a 5-year experience (1993–1998). J Clin Microbiol. 1999;37:1899–905.PubMedGoogle Scholar
- Gouriet F, Fenollar F, Patrice JY, Dancourt M, Raoult D. Use of shell-vial cell culture assay for isolation of bacteria from clinical specimens: 13 years of experience. J Clin Microbiol. 2005;43:4993–5002. DOIPubMedGoogle Scholar
- Maggi RG, Harms CA, Hohn AA, Pabst DA, McLellan WA, Walton WJ, Bartonella henselae in porpoise blood. Emerg Infect Dis. 2005;11:1894–8.PubMedGoogle Scholar
- Breitschwerdt EB, Maggi RG, Sigmon B, Nicholson WL. Isolation of Bartonella quintana from a woman and a cat following putative bite transmission. J Clin Microbiol. 2007;45:270–2. DOIPubMedGoogle Scholar
- Maggi RG, Breitschwerdt EB. Potential limitations of the 16S–23S rRNA intergenic region for the molecular detection of Bartonella species. J Clin Microbiol. 2005;43:1171–6. DOIPubMedGoogle Scholar
- Maggi RG, Chomel B, Hegarty BC, Henn J, Breitschwerdt EB. A Bartonella vinsonii berkhoffii typing scheme based upon 16S–23S ITS and Pap31 sequences from dog, coyote, gray fox, and human isolates. Mol Cell Probes. 2006;20:128–34. DOIPubMedGoogle Scholar
- Dalton MJ, Robinson LE, Copper J, Regnery RL, Olson JG, Childs JE. Use of Bartonella antigens for serologic diagnosis of cat-scratch disease at a national referral center. Arch Intern Med. 1995;155:1670–6. DOIPubMedGoogle Scholar
- Brouqui P, La Scola B, Roux V, Raoult D. Chronic Bartonella quintana bacteremia in homeless patients. N Engl J Med. 1999;340:184–9. DOIPubMedGoogle Scholar
- Arvand M, Schad SG. Isolation of Bartonella henselae DNA from the peripheral blood of a patient with cat scratch disease up to 4 months after the cat scratch injury. J Clin Microbiol. 2006;44:2288–90. DOIPubMedGoogle Scholar
- Kordick DL, Brown TT, Shin KO, Breitschwerdt EB. Clinical and pathological evaluation of chronic Bartonella henselae or Bartonella clarridgeiae infection in cats. J Clin Microbiol. 1999;37:1536–47.PubMedGoogle Scholar
- Arvand M, Schubert H, Viezens J. Emergence of distinct genetic variants in the population of primary Bartonella henselae isolates. Microbes Infect. 2006;8:1315–20. DOIPubMedGoogle Scholar
- Adelson ME, Rao RV, Tilton RC, Cabets K, Eskow E, Fein L, Prevalence of Borrelia burgdorferi, Bartonella spp., Babesia microti, and Anaplasma phagocytophila in Ixodes scapularis ticks collected in Northern New Jersey. J Clin Microbiol. 2004;42:2799–801. DOIPubMedGoogle Scholar
- Giladi M, Maman E, Paran D, Bickels J, Comaneshter D, Avidor B, Cat-scratch disease-associated arthropathy. Arthritis Rheum. 2005;52:3611–7. DOIPubMedGoogle Scholar
Figures
Table
Cite This ArticleTable of Contents – Volume 13, Number 6—June 2007
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Edward B. Breitschwerdt, North Carolina State University College of Veterinary Medicine, 4700 Hillsborough St, Raleigh, NC 27606, USA; .
Top