Volume 13, Number 7—July 2007
Perspective
Brazilian Vaccinia Viruses and Their Origins
Figure 3
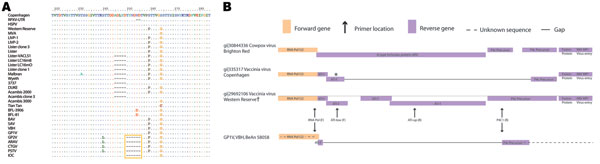
Figure 3. A) Partial amino acid alignment of A56R vaccinia virus Copenhagen (VACV-COP). Sequences were retrieved from GenBank and aligned using the BioEdit program (www.mbio.ncsu.edu/bioedit/bioedit.html). VACV-COP (M35027, 161908-161957) is used as the reference sequence; International Union of Pure and Applied Chemistry symbols, amino acids; dots, residues identical to the reference sequence; dashes, gaps created by the alignment; gold box surrounds the 18-nt deletion once proposed as a Brazilian (BRZ)-VACV molecular identifier. RPXV-UTR, rabbitpox virus-Utrecht; HSPV, horsepox virus; MVA, modified vaccinia Ankara; LIVP, Liverpool; BFL, buffalopox; BAV, BeAn 58058 virus; SAV, SpAn232 virus; VBH, Belo Horizonte virus; GP, Guarani virus; ARAV, Araçatuba virus; CTGV, Cantagalo virus; PSTV, Passatempo virus; IOC, Institute Oswaldo Cruz. B) Depiction of the rpo132-ATI-p4c-A27 region in Cowpox virus Brighton Red (CPXV-BR) and VACV strains. Positions of primers RNApol, ATI-low, ATI-up, and P4c1 are illustrated. These primers have been used for the molecular diagnosis of Brazilian (BRZ)-VACV (5,26). The 300-bp fragment amplified by primers RNApol and P4c1 from GP1V and VBH was sequenced by Trindade et al. (9,13). Sequence outside this region is unknown. ATI-f is used to designate coding regions that resemble fragments of the wild-type (CPXV-BR) ATI gene. The gap in VACV-COP lies within the A26L gene. *This gene is often referred to as ATI due to its original description. However, the coding region is actually a fusion of the C-terminus of ATI and the N-terminus of P4c where the sequence in between has been deleted. The gene is currently identified by the Poxviridae Bioinformatics Resource (http://athena.bioc.uvic.ca/database.php?db=poxviridae) as belonging to the P4c precursor family. †Western Reserve is presumed similar in structure to SpAn232 virus, GP2V, ARAV, and PSTV due to similar restriction fragment length polymorphism results (6,7,11,13).
References
- Lewis-Jones S. Zoonotic poxvirus infections in humans.Curr Opin Infect Dis. 2004;17:81–9. DOIPubMedGoogle Scholar
- Fernandes T. The smallpox vaccine: its first century in Brazil (from the Jennerian to the animal vaccine)[in Spanish]. Hist Cienc Saude Manguinhos. 1999;6:29–51. DOIPubMedGoogle Scholar
- Fenner F, Henderson DA, Arita I, Jezek Z, Ladnyi ID. Smallpox and its eradication. Geneva: World Health Organization; 1988.
- Fonseca FG, Lanna MC, Campos MAS, Kitajima EW, Peres JN, Golgher RR, Morphological and molecular characterization of the poxvirus BeAn 58058.Arch Virol. 1998;143:1171–86. DOIPubMedGoogle Scholar
- Marques JT, Trindade GS, da Fonseca FG, dos Santos JR, Bonjardim CA, Ferreira PCP, Characterization of ATI, TK and IFN-alpha/betaR genes in the genome of the BeAn 58058 virus, a naturally attenuated wild orthopoxvirus.Virus Genes. 2001;23:291–301. DOIPubMedGoogle Scholar
- da Fonseca FG, Trindade GS, Silva RL, Bonjardim CA, Ferreira PC, Kroon EG. Characterization of a vaccinia-like virus isolated in a Brazilian forest.J Gen Virol. 2002;83:223–8.PubMedGoogle Scholar
- de Souza Trindade G, da Fonseca FG, Marques JT, Nogueira ML, Mendes LC, Borges AS, Araçatuba virus: a vaccinialike virus associated with infection in humans and cattle.Emerg Infect Dis. 2003;9:155–60.PubMedGoogle Scholar
- Damaso CR, Esposito JJ, Condit RC, Moussatche N. An emergent poxvirus from humans and cattle in Rio de Janeiro State: Cantagalo virus may derive from Brazilian smallpox vaccine.Virology. 2000;277:439–49. DOIPubMedGoogle Scholar
- Trindade GS, da Fonseca FG, Marques JT, Diniz S, Leite JA, De Bodt S, Belo Horizonte virus: a vaccinia-like virus lacking the A-type inclusion body gene isolated from infected mice.J Gen Virol. 2004;85:2015–21. DOIPubMedGoogle Scholar
- Nagasse-Sugahara TK, Kisielius JJ, Ueda-Ito M, Curti SP, Figueiredo CA, Cruz AS, Human vaccinia-like virus outbreaks in São Paulo and Goias States, Brazil: virus detection, isolation and identification.Rev Inst Med Trop Sao Paulo. 2004;46:315–22. DOIPubMedGoogle Scholar
- Leite JA, Drumond BP, Trindade G, Lobato ZIP, da Fonseca FG, Santos JR, Passatempo virus, a vaccinia virus strain, Brazil.Emerg Infect Dis. 2005;11:1935–8.PubMedGoogle Scholar
- Lobato ZIP, Trindade GS, Frois MCM, Ribeiro EBT, Dias GRC, Teixeira BM, Outbreak of exanthemal disease caused by vaccinia virus in human and cattle in Zona da Mata region, Minas Gerais[in Portuguese]. Arq Bras Med Vet Zootec.2005;57:423–9. DOIGoogle Scholar
- Trindade GS, Lobato ZIP, Drumond BP, Leite JA, Trigueiro RC, Guedes MIMC, Isolation of two vaccinia virus strains from a single bovine vaccinia outbreak in rural area from Brazil: implications on the emergence of zoonotic orthopoxviruses.Am J Trop Med Hyg. 2006;75:486–90.PubMedGoogle Scholar
- Johnson GP, Goebel SJ, Paoletti E. An update on the vaccinia virus genome.Virology. 1993;196:381–401. DOIPubMedGoogle Scholar
- Dumbell K, Richardson M. Virological investigations of specimens from buffaloes affected by buffalopox in Maharashtra State, India between 1985 and 1987.Arch Virol. 1993;128:257–67. DOIPubMedGoogle Scholar
- Kolhapure RM, Tupe CD, Deolankar RP, Raut CG, Basu A, Dama BM, Investigation of buffalopox outbreaks in Maharashtra State during 1992-1996.Indian J Med Res. 1997;106:441–6.PubMedGoogle Scholar
- Singh RK, Hosamani M, Balamurugan V, Satheesh CC, Rasool TJ, Yadav MP. Comparative sequence analysis of envelope protein genes of Indian buffalopox virus isolates.Arch Virol. 2006;151:1995–2005. DOIPubMedGoogle Scholar
- Tulman ER, Delhon G, Afonso CL, Lu Z, Zsak L, Sandybaev NT, Genome of horsepox virus.J Virol. 2006;80:9244–58. DOIPubMedGoogle Scholar
- Swofford DL. PAUP*. Phylogenetic Analysis Using Parsimony (*and other methods). Version 4. Sunderland (MA): Sinauer Associates; 2003.
- Huelsenbeck JP, Ronquist F. MRBAYES: Bayesian inference of phylogenetic trees.Bioinformatics. 2001;17:754–5. DOIPubMedGoogle Scholar
- Ronquist F, Huelsenbeck JP. MrBayes 3: Bayesian phylogenetic inference under mixed models.Bioinformatics. 2003;19:1572–4. DOIPubMedGoogle Scholar
- Posada D, Crandall KA. MODELTEST: testing the model of DNA substitution.Bioinformatics. 1998;14:817–8. DOIPubMedGoogle Scholar
- Posada D, Buckley TR. Model selection and model averaging in phylogenetics: advantages of Akaike information criterion and Bayesian approaches over likelihood ratio tests.Syst Biol. 2004;53:793–808. DOIPubMedGoogle Scholar
- Ropp SL, Jin Q, Knight JC, Massung RF, Esposito JJ. PCR strategy for identification and differentiation of smallpox and other orthopoxviruses.J Clin Microbiol. 1995;33:2069–76.PubMedGoogle Scholar
- Damaso CR, Reis SA, Jesus DM, Lima PS, Moussatche N. A PCR-based assay for detection of emerging vaccinia-like viruses isolated in Brazil.Diagn Microbiol Infect Dis. 2006;57:39–46. DOIPubMedGoogle Scholar
- Meyer H, Damon IK, Esposito JJ. Orthopoxvirus diagnostics. In: Isaacs, SN, editor. Vaccinia virus and poxvirology: methods and protocols. Totowa (NJ): Humana Press; 2004 p. 119–33.
- Lewis A, Bok K, Perez O, De Fillipo J, Paolazzi C, Gomes JA. Characterization of a Vaccinia virus strain used to produce smallpox vaccine in Argentina between 1937 and 1970.Arch Virol. 2005;150:1485–91. DOIPubMedGoogle Scholar
- World Health Organization. Archives of the Smallpox Eradication Programme. [cited 2007 May 17.] Available from http://www.who.int/archives/fonds_collections/bytitle/fonds_6/en/index.html
- Turner GS. Respiratory infection of mice with vaccinia virus.J Gen Virol. 1967;1:399–402. DOIPubMedGoogle Scholar
- Wokatsch R. Vaccinia virus. In: Majer M, Plotkin SA, editors. Strains of human viruses. Basel: S. Karger; 1972.
- de Souza Lopes O, Lacerda JP, Fonseca IE, Castro DP, Forattini OP, Rabello EX. Cotia virus: a new agent isolated from sentinel mice in São Paulo, Brazil.Am J Trop Med Hyg. 1965;14:156–7.PubMedGoogle Scholar
- Ueda Y, Dumbell KR, Tsuruhara T, Tagaya I. Studies on Cotia virus—an unclassified poxvirus.J Gen Virol. 1978;40:263–76. DOIPubMedGoogle Scholar
- Ueda Y, Morikawa S, Watanabe T. Unclassified poxvirus: characterization and physical mapping of Cotia virus DNA and location of a sequence capable of encoding a thymidine kinase.Virology. 1995;210:67–72. DOIPubMedGoogle Scholar
- Esposito JJ, Palmer EL, Borden EC, Harrison AK, Obijeski JF, Murphy FA. Studies on the poxvirus Cotia.J Gen Virol. 1980;47:37–46. DOIPubMedGoogle Scholar
- Conservational International [cited 2006 Sep 2]. Available from http://www.biodiversityhotspots.org/xp/Hotspots/atlantic_forest
- Benchimol JL. Febre Amarela: a doença e a vacina, uma historia inacabada. Rio de Janeiro: Editora Fiocruz; 2001.