Nosocomial Outbreak of Novel Arenavirus Infection, Southern Africa
Janusz T. Paweska, Nivesh H. Sewlall, Thomas G. Ksiazek, Lucille H. Blumberg, Martin J. Hale, W. Ian Lipkin, Jacqueline Weyer, Stuart T. Nichol, Pierre E. Rollin, Laura K. McMullan, Christopher D. Paddock, Thomas Briese, Joy Mnyaluza, Thu-Ha Dinh, Victor Mukonka, Pamela Ching, Adriano Duse, Guy Richards, Gillian de Jong, Stefano Tempia, Bridget Ikalafeng, Charles Mugero, Chika Asomugha, Mirriam M. Malotle, Dorothy M. Nteo, Eunice Misiani, Robert Swanepoel

, Sherif R. Zaki, Investigation Teams
1, and members of the Outbreak Control
Author affiliations: National Institute for Communicable Diseases, Sandringham, South Africa (J.T. Paweska, L.H. Blumberg, J. Weyer, G de Jong, C. Cohen, R. Swanepoel); University of the Witwatersrand, Johannesburg, South Africa (N.H. Sewlall, M.J. Hale, A. Duse); Centers for Disease Control and Prevention (CDC), Atlanta, Georgia, USA (T.G. Ksiazek, S.T. Nichol, P.E. Rollin, L.K. McMullan, C.D. Paddock, T.-H. Dinh, S.R. Zaki); Columbia University, New York, New York, USA (W.I. Lipkin, T. Briese); Gauteng Department of Health, Johannesburg (J. Mnyaluza, B. Ikalafeng, C. Asomugha); CDC Global AIDS Program, Pretoria, South Africa (T.-H. Dinh); Ministry of Health, Lusaka, Zambia (V. Mukonka); CDC Global AIDS Program, Lusaka (P. Ching); Charlotte Maxeke Hospital, Johannesburg (G. Richards); National Department of Health, Pretoria (C. Mugero, E. Misiani); Field Epidemiology and Laboratory Training Programme, Johannesburg (M.M. Malotle, D.M. Nteo); University of Texas Medical Branch, Galveston, Texas, USA (T.G. Ksiasek)
Main Article
Figure 3
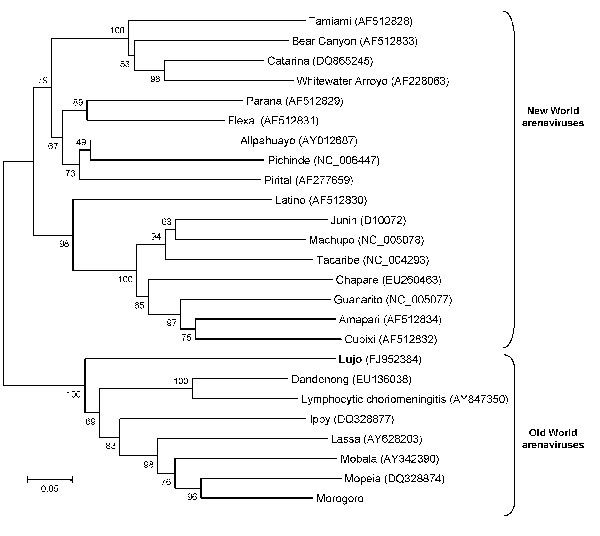
Figure 3. Neighbor-joining tree reconstructed by using bootstrap analysis with 1,000 pseudoreplicate datasets showing the phylogenetic relationship of known arenaviruses (data derived from GenBank) to the novel Lujo arenavirus from southern Africa (boldface), inferred from a 619-nt region of the 5′ end of the nucleoprotein gene. GenBank accession numbers for nucleotide sequence data are shown on the tree. Scale bar indicates 5% divergence.
Main Article
Page created: December 08, 2010
Page updated: December 08, 2010
Page reviewed: December 08, 2010
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.
