Volume 15, Number 3—March 2009
Research
Sources of Hepatitis E Virus Genotype 3 in the Netherlands
Figure 2
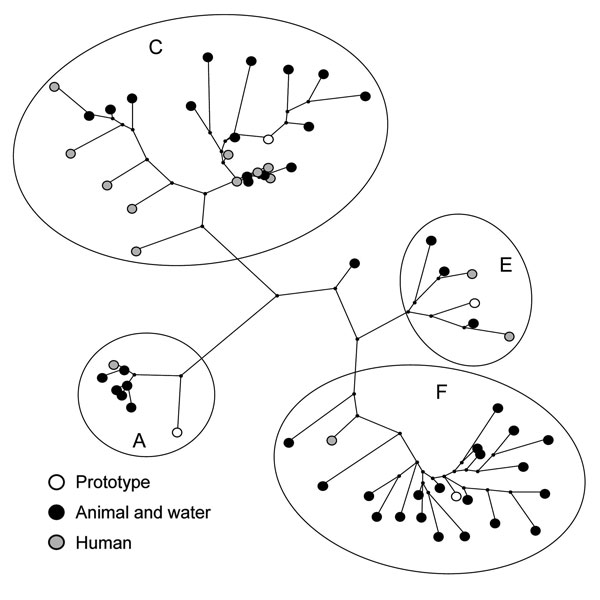
Figure 2. Maximum-parsimony tree of hepatitis E virus (HEV) sequences detected in environmental samples and patients during 2004–2006, based on a 148-nt sequence of open reading frame 2 (nt 6322–6469 of strain M73218). Origins of HEV sequences and genotype 3 clusters are indicated. Sequences are compared with prototype sequences of different clusters of HEV genotype 3. Prototypes correspond with the following GenBank accession nos.: A) US1, AF060668; C) NLSW105, AF336298; E) UK-swine p354: AF503511; F) G1, AF110391. The following accession numbers have been used for phylogenetic analysis of human isolates: AB385844–AB385848, AB385850–AB385852, DQ200279, DC200282–DQ200284, DQ200287, DQ200289, DQ200292, DQ200293. Accession numbers of animal and environmental samples are those in Figure 1.