Volume 15, Number 4—April 2009
Letter
Leptospira noguchii and Human and Animal Leptospirosis, Southern Brazil
Figure
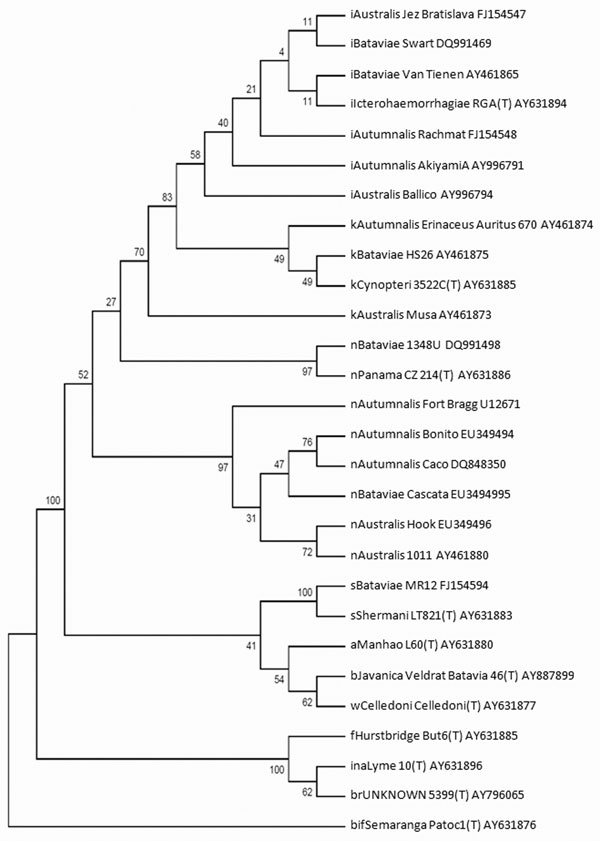
Figure. Dendogram constructed by using the neighbor-joining algorithm, based on a 1,180-bp sequence of the 16S rRNA gene demonstrating the position of the Brazilian strains (Bonito, Cascata, and Hook) within the Leptospira noguchii species. This dendogram summarizes, by bootstrap-based topology, the evolutionary relationship among L. noguchii strains. The bootstrap consensus values are indicated over each root. The initial lowercase letters indicate the respective species to which each strain belongs: i, L. interrogans; k, L. kirschneri; n, L. noguchii; b, L. borgpetersenii; w, L. weilii; s, L. santarosai; a, L. alexanderi; f, L. fainei; in, L. inadai; br, L. broomi; bif, L. biflexa. (T) indicates the type-strain for each species. The GenBank accession number follows the strain identification.