Volume 15, Number 6—June 2009
Research
Lineage 2 West Nile Virus as Cause of Fatal Neurologic Disease in Horses, South Africa
Figure 3
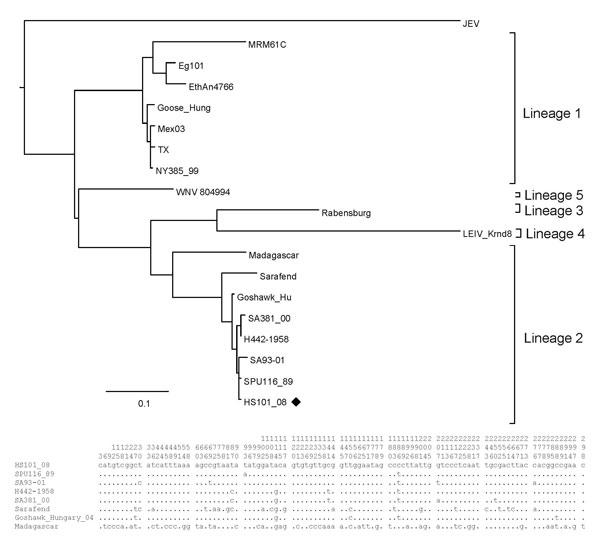
Figure 3. Maximum-likelihood analysis of the E-protein region of a West Nile virus isolate, HS101_08 (black diamond), recovered from horses in 2008 compared with isolates obtained from humans and animals from South Africa and other regions of the world. Nucleotide differences between lineage 2 strains included in the alignment are shown in the summarized alignment below the tree, indicating only unique nucleotides. Vertical numbers above the alignment indicate the position of each variable site on the gene fragment. Scale bar indicates nucleotide substitutions per site.
Page created: December 08, 2010
Page updated: December 08, 2010
Page reviewed: December 08, 2010
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.