Volume 15, Number 9—September 2009
Research
Predicting Phenotype and Emerging Strains among Chlamydia trachomatis Infections
Figure 1
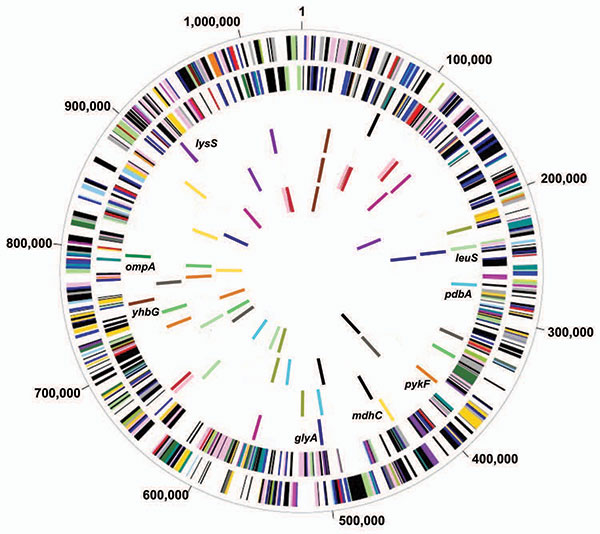
Figure 1. Comparison of 14 housekeeping genes among genome sequences of 4 Chlamydiaceae species and 7 strains. Circle 1, genes on forward Chlamydia trachomatis strand, color coded by role category; Circle 2, genes on reverse C. trachomatis strand; Circle 3, multilocus sequence typing (MLST) candidates, C. trachomatis; Circle 4, MLST candidates, C. pneumoniae AR39; Circle 5, MLST candidates, C. caviae (GPIC); Circle 6, MLST candidates, C. muridarum (MoPn). Colors in circles 3, 4, 5 and 6 are consistent for each gene across genomes i.e., “blue” gene in each circle is ortholog in that genome for “blue” gene in C. trachomatis. Blue, glyA, serine hydroxymethyl-transferase; red, tryptophanyl-tRNA synthetase; yellow, mdhC, malate dehydrogenase; green, V-type ATPase, subunit A; cyan, pdhA, pyruvate dehydrogenase; black, GTP-binding protein lepa; magenta, transcription termination factor rho; brown, yhbG, probable ABC transporter ATP-binding protein; orange, pykF, pyruvate kinase; olive green, conserved hypothetical protein; gray, acetyl-CoA carboxylase beta subunit; pink, threonyl-tRNA synthetase; violet, lysS, lysyl-tRNA synthetase; light green, leuS, leucyl-tRNA synthetase. Those denoted in boldface above were used for C. trachomatis MLST. ompA gene location is shown for C. trachomatis (dark green).