Volume 17, Number 1—January 2011
Research
Endurance, Refuge, and Reemergence of Dengue Virus Type 2, Puerto Rico, 1986–2007
Figure 2
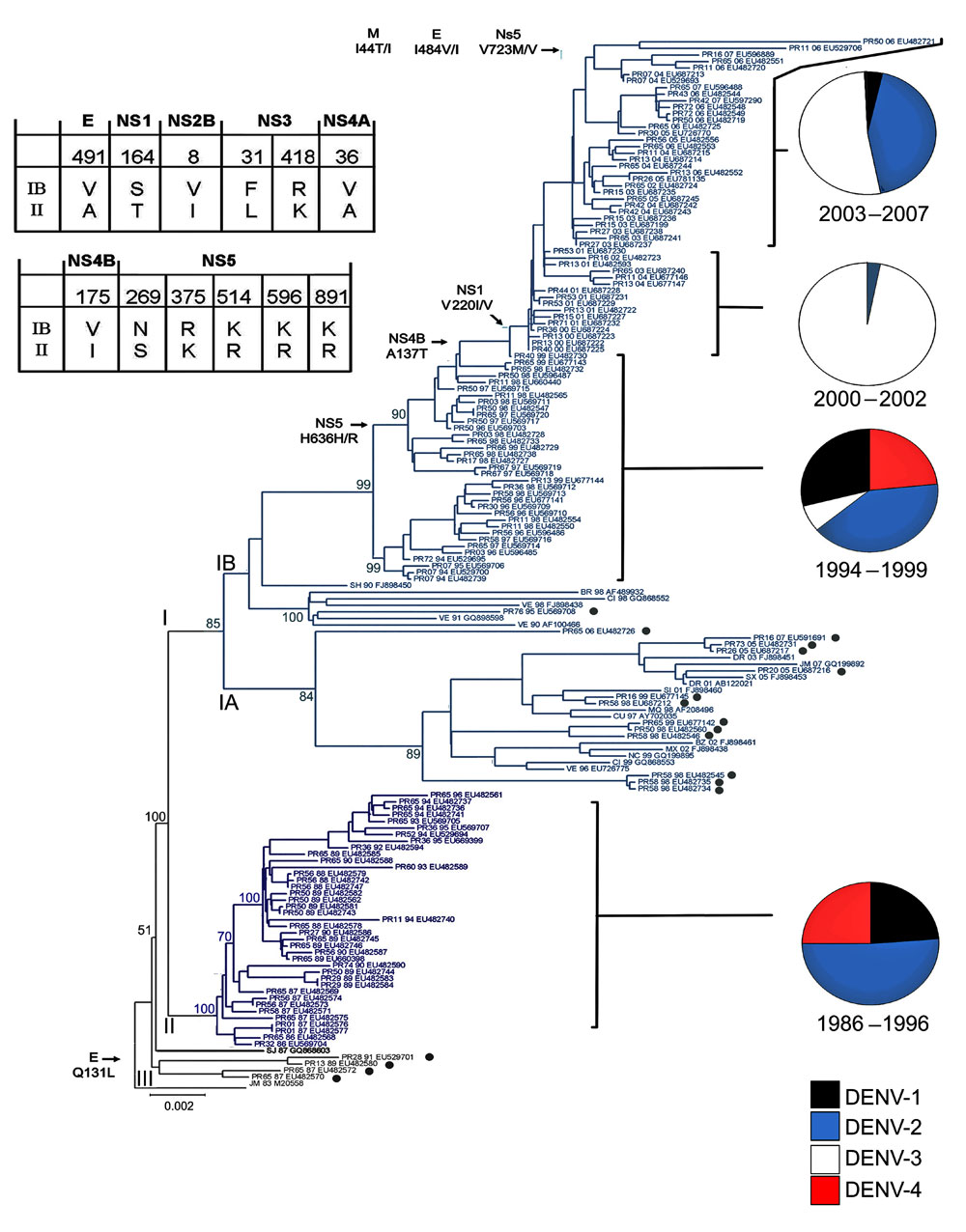
Figure 2. Evolution of dengue virus (DENV) serotype 2, Puerto Rico. Maximum likelihood phylogeny of the 140 Puerto Rico and 20 international isolates of DENV-2 (see number of isolates by year below). Names of clades (I, II, and III) and subclades (IA, IB) are shown at the base of their respective branches on the phylogeny tree. Clade II (dark blue) circulated during 1986–1996 and clade I (light blue) during 1994–2007. Subclade IA and clade III represent foreign, transient reintroductions throughout the 22-year study period. Black dots indicate 18 isolates from Puerto Rico with phylogenetic associations closer to foreign isolates than to other Puerto Rico viruses. Two-letter geographic codes indicate origin (PR, Puerto Rico; JM, Jamaica; SJ, Saint John; SH, Saint Thomas; VE, Venezuela; Cl, Colombia; NC, Nicaragua; MX, Mexico; CU, Cuba; SI, Saint Kits; DR, Dominican Republic; SX, Saint Croix; BR, Brazil). PR code is accompanied by a 2-digit number that indicates geographic location (municipality) on the island. The 2-digit numbers between the geographic codes and the GenBank accession numbers indicate the year of the corresponding case of each isolate. Bootstrap values are shown for all clades (I, II, and II), subclades (IA, IB), and most immediate lineages. Twelve amino acid changes associated with IB/II differences are shown on the Table; 6 other selected changes across subclade IB are shown on the left at the base of the branches exhibiting the respective changes. Amino acid numbers refer to the position in each DENV protein. Relevant epidemiologic events are highlighted to the right with pie charts showing the relative levels of each serotype isolated for that period Black, DENV-1; blue, DENV-2; white, DENV-3; red, DENV-4. Scale bar indicates nucleotide substitutions per site.
1These authors contributed equally to this article.