Dynamics of Cholera Outbreaks in Great Lakes Region of Africa, 1978–2008
Didier Bompangue Nkoko, Patrick Giraudoux, Pierre-Denis Plisnier, Annie Mutombo Tinda, Martine Piarroux, Bertrand Sudre, Stephanie Horion, Jean-Jacques Muyembe Tamfum, Benoît Kebela Ilunga, and Renaud Piarroux

Author affiliations: Université de Franche-Comté, Besançon, France (D. Bompangue Nkoko, P. Giraudoux, M. Piarroux, B. Sudre); Ministère de la Santé Publique, Kinshasa, Democratic Republic of Congo (D. Bompangue Nkoko, A. Mutombo Tinda, J.-J. Muyembe Tamfum, B. Kebela Ilunga); Royal Museum for Central Africa, Tervuren, Belgium (P.-D. Plisnier); Joint Research Centre of the European Commission, Ispra, Italy (S. Horion); Université de Kinshasa, Kinshasha (J.-J. Muyembe Tamfum); Université de la Méditerranée, Marseille, France (R. Piarroux); University Hospital La Timone, Marseille (R. Piarroux)
Main Article
Figure 3
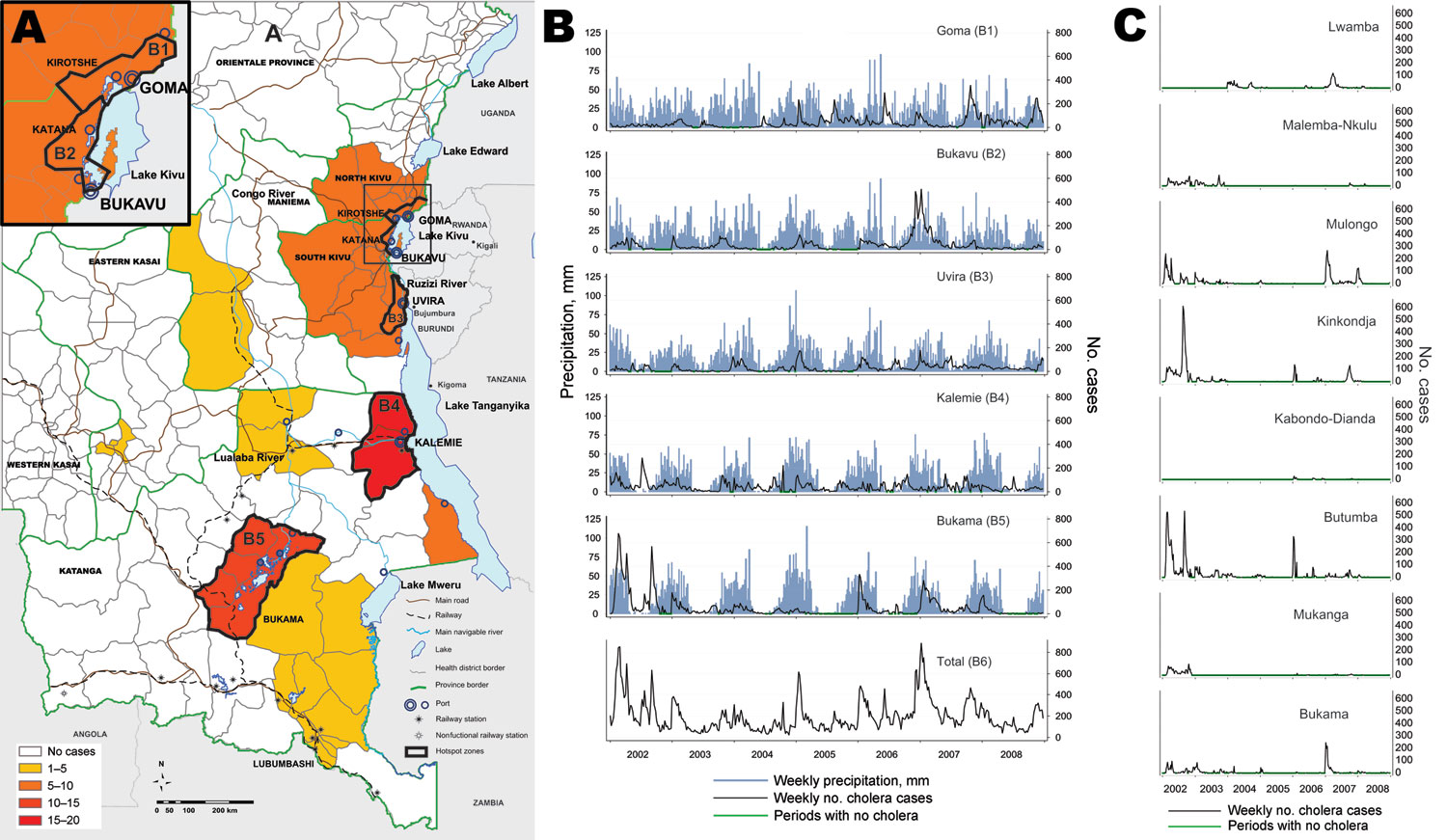
Figure 3. Temporal-spatial evolution of cholera cases in 5 hotspots in the African Great Lakes region, 2002–2008. A) Spatial distribution of cholera in the provinces of Katanga, North Kivu, and South Kivu (Democratic Republic of Congo). Health districts are colored according to the risk ratio of the cluster, as calculated by using SatSCan software (Kulldorf, Cambridge, UK). B) Evolution of the weekly number of cholera cases in the 5 hotspots (B1–B5). B1) Goma and Kirotshe health districts; B2) Bukavu and Katana health districts; B3) Uvira health district; B4) Kalemie and Nyemba health districts; B5) 8 health districts in the Upper Congo River Basin (see district names in panel C); B6) total cases for the 5 hotspots. Green indicates periods without cholera; blue indicates estimated weekly rainfalls. The global curve did not show any remission periods. C) Evolution of the weekly number of cholera cases in the 8 health districts composing the Upper Congo Basin hotspot. The epidemic curve in B5 was composed of partially synchronous epidemics separated by periods of lull.
Main Article
Page created: October 19, 2011
Page updated: October 19, 2011
Page reviewed: October 19, 2011
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.
