Volume 17, Number 2—February 2011
Letter
Clonal Spread of Streptococcus pyogenes emm44 among Homeless Persons, Rennes, France
Figure
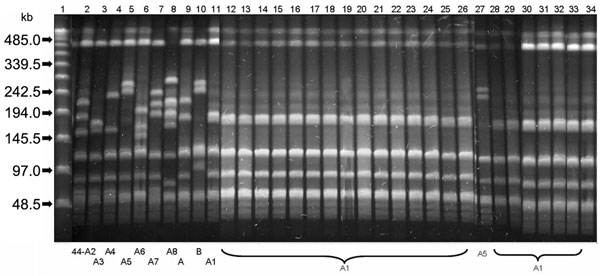
Figure. Pulsed-field gel electrophoresis (PFGE) patterns of SmaI-restricted chromosomal DNA of Streptococcus pyogenes emm44 strains. Lane 1, Bacteriophage Lambda ladder PFGE Marker (New England Biolabs Inc., Beverly, MA, USA); lanes 2–11, PFGE patterns 44-A2, 44-A3, 44-A4, 44-A5, 44-A6, 44-A7, 44-A8, 44-A, 44-B, and 44-A1 of emm44 unrelated control strains; lanes 12–26 and 28–34, 22 identical 44-A1 PFGE patterns shared by the tetracycline-resistant outbreak isolates; lane 27, PFGE pattern 44-A5 of the nonclonal emm44 strain isolated during the same outbreak, which differs by 4 bands from the pattern 44-A1.
1These authors contributed equally to this article.
Page created: July 13, 2011
Page updated: July 13, 2011
Page reviewed: July 13, 2011
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.