Volume 17, Number 8—August 2011
Dispatch
Atypical Pestivirus and Severe Respiratory Disease in Calves, Europe
Figure
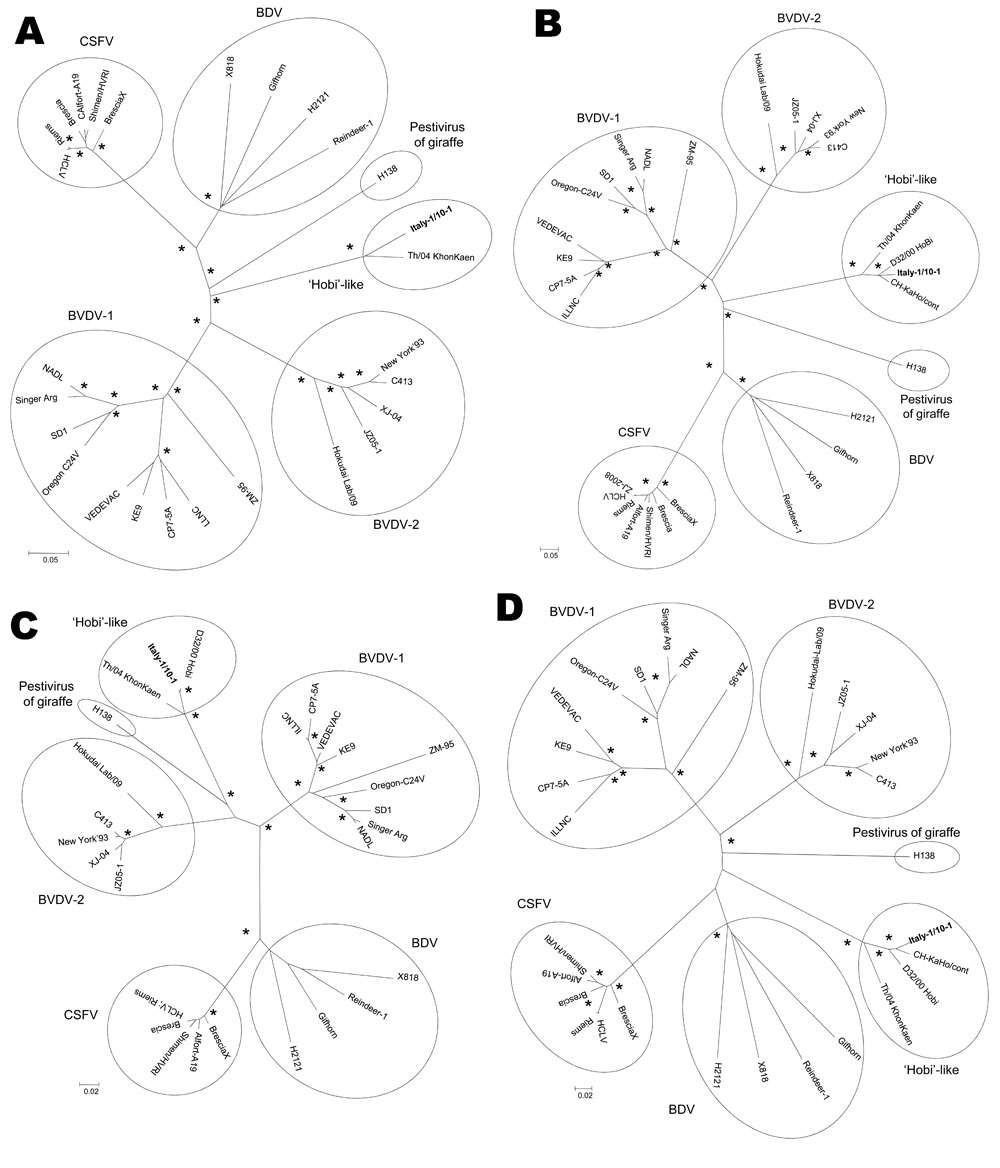
Figure. Neighbor-joining unrooted trees based on the full-length genome (A), E2 (B), 5′ untranslated region (C), and Npro (D) sequences of members of the genus Pestivirus. For phylogenetic tree construction, pestivirus sequences listed in the Table were used. Asterisks indicate strong statistical support for a node by a bootstrap value of 75%–100%. Scale bars represent estimated numbers of nucleotide substitutions per site. CSFV, classical swine fever virus; BVDV-1, bovine viral diarrhea virus type 1; BVDV-2, bovine viral diarrhea virus type 2; BDV, border disease virus.
Page created: August 15, 2011
Page updated: August 15, 2011
Page reviewed: August 15, 2011
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.