Volume 18, Number 9—September 2012
Dispatch
Multiple Synchronous Outbreaks of Puumala Virus, Germany, 2010
Figure 2
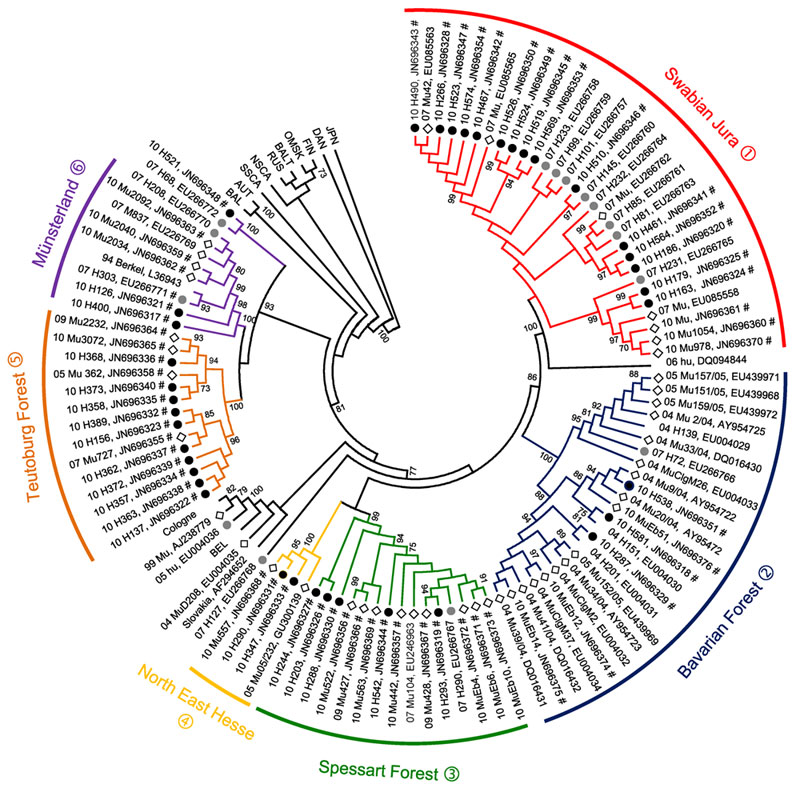
Figure 2. . . . Neighbor-joining phylogenetic tree (TN93 evolutionary model) of Puumala virus (PUUV) strains constructed on the basis of partial sequences of the small segment (504-nt sequence, nt positions 392–894). Bootstrap values >70%, calculated from 10,000 replicates, are shown at the tree branches. Analysis was performed by using MEGA5 software (www.megasoftware.net). PUUV-like sequences from Japan (JPN) were used as outgroup. Numbers from 04 to 11 in front of the sample names indicate the year (2004–2011) when the sample was collected. Black dots indicate human samples from 2010, gray dots indicate human samples from 2007, and diamonds indicate rodent samples. Novel sequences from this study are indicated by the symbol #. For numbers (1–6) of PUUV clades that correspond to the 6 defined outbreak regions, see Figure 1. Phylogenetic clades are shown in parentheses followed by names and numbers. For clarity, previously characterized PUUV clades from other parts of Europe are shown in simplified form. BEL, Belgium; BAL, Balkan; AUT, Austrian; SSCA, South Scandinavian; NSCA, North Scandinavian; RUS, Russian; BALT, Baltic; OMSK, Russian from Omsk region; FIN, Finnish; DAN, Danish.
1These authors contributed equally to this article.