Volume 19, Number 8—August 2013
Research
Aichi Virus in Sewage and Surface Water, the Netherlands
Figure 3
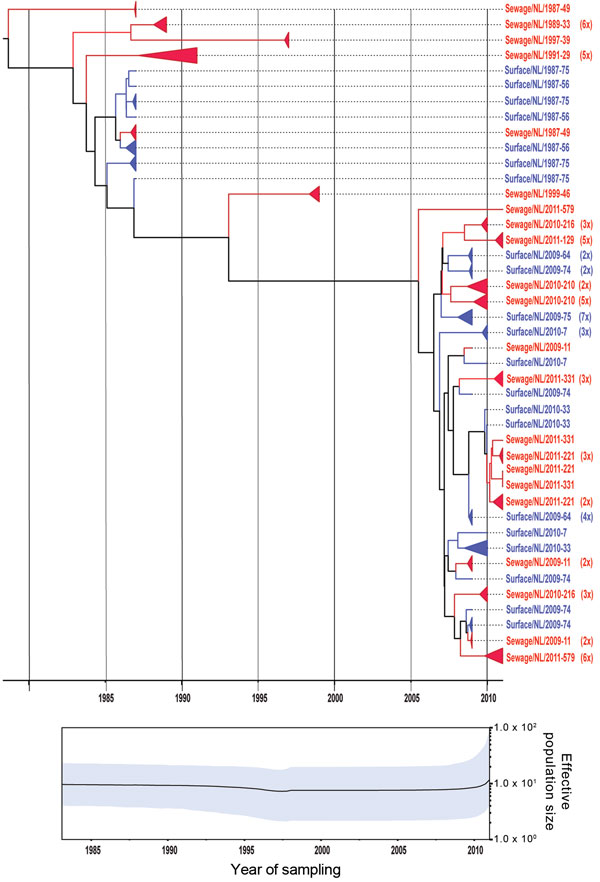
Figure 3. . . Phylogenetic relationships and genetic diversity over time for 166 sequences of Aichi virus genotype B strains collected in the Netherlands. A) Maximum-clade credibility tree was generated by the Bayesian Markov chain Monte Carlo method in BEAST (28), based on a multiple alignment of nucleotide sequences (481-nt) of the viral protein 1 region. The tree is rooted to the most recent common ancestor, visualized in FigTree (http://tree.bio.ed.ac.uk/software/figtree/), and plotted on a temporal y-axis scale using the sampling dates. Aichi virus strains from the Netherlands isolated from sewage (red) and surface waters (blue) are indicated. Clusters of sequences of the same sample are represented by triangles (a collapsed branch), and the number of isolates in each triangle is shown in parentheses. B) Bayesian skyline plot obtained by analyzing different Aichi virus sequences sampled at different times. The results are a relative measure for genetic diversity through time. The line represents the median, and the shaded area represents the 95% highest posterior density of the number of isolates.
References
- Clasen T, Schmidt WP, Rabie T, Roberts I, Cairncross S. Interventions to improve water quality for preventing diarrhoea: systematic review and meta-analysis. BMJ. 2007;334:782. DOIPubMedGoogle Scholar
- Tapparel C, Siegrist F, Petty TJ, Kaiser L. Picornavirus and enterovirus diversity with associated human diseases. Infect Genet Evol. 2013;14:282–93. DOIPubMedGoogle Scholar
- Yamashita T, Kobayashi S, Sakae K, Nakata S, Chiba S, Ishihara Y, Isolation of cytopathic small round viruses with BS-C-1 cells from patients with gastroenteritis. J Infect Dis. 1991;164:954–7. DOIPubMedGoogle Scholar
- Yamashita T, Sakae K, Tsuzuki H, Suzuki Y, Ishikawa N, Takeda N, Complete nucleotide sequence and genetic organization of Aichi virus, a distinct member of the Picornaviridae associated with acute gastroenteritis in humans. J Virol. 1998;72:8408–12 .PubMedGoogle Scholar
- Yamashita T, Ito M, Tsuzuki H, Sakae K. Identification of Aichi virus infection by measurement of immunoglobulin responses in an enzyme-linked immunosorbent assay. J Clin Microbiol. 2001;39:4178–80. DOIPubMedGoogle Scholar
- Oh DY, Silva PA, Hauroeder B, Diedrich S, Cardoso DD, Schreier E. Molecular characterization of the first Aichi viruses isolated in Europe and in South America. Arch Virol. 2006;151:1199–206. DOIPubMedGoogle Scholar
- Pham NT, Khamrin P, Nguyen TA, Kanti DS, Phan TG, Okitsu S, Isolation and molecular characterization of Aichi viruses from fecal specimens collected in Japan, Bangladesh, Thailand, and Vietnam. J Clin Microbiol. 2007;45:2287–8. DOIPubMedGoogle Scholar
- Yang S, Zhang W, Shen Q, Yang Z, Zhu J, Cui L, Aichi virus strains in children with gastroenteritis, China. Emerg Infect Dis. 2009;15:1703–5. DOIPubMedGoogle Scholar
- Ambert-Balay K, Lorrot M, Bon F, Giraudon H, Kaplon J, Wolfer M, Prevalence and genetic diversity of Aichi virus strains in stool samples from community and hospitalized patients. J Clin Microbiol. 2008;46:1252–8. DOIPubMedGoogle Scholar
- Jonsson N, Wahlstrom K, Svensson L, Serrander L, Lindberg AM. Aichi virus infection in elderly people in Sweden. Arch Virol. 2012;157:1365–9. DOIPubMedGoogle Scholar
- Kaikkonen S, Rasanen S, Ramet M, Vesikari T. Aichi virus infection in children with acute gastroenteritis in Finland. Epidemiol Infect. 2010;138:1166–71. DOIPubMedGoogle Scholar
- Reuter G, Boldizsar A, Papp G, Pankovics P. Detection of Aichi virus shedding in a child with enteric and extraintestinal symptoms in Hungary. Arch Virol. 2009;154:1529–32. DOIPubMedGoogle Scholar
- Sdiri-Loulizi K, Hassine M, Gharbi-Khelifi H, Sakly N, Chouchane S, Guediche MN, Detection and genomic characterization of Aichi viruses in stool samples from children in Monastir, Tunisia. J Clin Microbiol. 2009;47:2275–8. DOIPubMedGoogle Scholar
- Reuter G, Boros A, Pankovics P. Kobuviruses—a comprehensive review. Rev Med Virol. 2011;21:32–41. DOIPubMedGoogle Scholar
- Yamashita T, Sugiyama M, Tsuzuki H, Sakae K, Suzuki Y, Miyazaki Y. Application of a reverse transcription–PCR for identification and differentiation of Aichi virus, a new member of the Picornavirus family associated with gastroenteritis in humans. J Clin Microbiol. 2000;38:2955–61 .PubMedGoogle Scholar
- Pham NT, Trinh QD, Nguyen TA, Dey SK, Phan TG. Hoang le P, et al. Development of genotype-specific primers for differentiation of genotypes A and B of Aichi viruses. J Virol Methods. 2009;156:107–10.
- Oberste MS, Maher K, Kilpatrick DR, Pallansch MA. Molecular evolution of the human enteroviruses: correlation of serotype with VP1 sequence and application to picornavirus classification. J Virol. 1999;73:1941–8 .PubMedGoogle Scholar
- Lodder WJ, de Roda Husman AM. Presence of noroviruses and other enteric viruses in sewage and surface waters in the Netherlands. Appl Environ Microbiol. 2005;71:1453–61. DOIPubMedGoogle Scholar
- Sdiri-Loulizi K, Hassine M, Aouni Z, Gharbi-Khelifi H, Sakly N, Chouchane S, First molecular detection of Aichi virus in sewage and shellfish samples in the Monastir region of Tunisia. Arch Virol. 2010;155:1509–13. DOIPubMedGoogle Scholar
- Alcalá A, Vizzi E, Rodriguez-Diaz J, Zambrano JL, Betancourt W, Liprandi F. Molecular detection and characterization of Aichi viruses in sewage-polluted waters of Venezuela. Appl Environ Microbiol. 2010;76:4113–5. DOIPubMedGoogle Scholar
- Kitajima M, Haramoto E, Phanuwan C, Katayama H. Prevalence and genetic diversity of Aichi viruses in wastewater and river water in Japan. Appl Environ Microbiol. 2011;77:2184–7. DOIPubMedGoogle Scholar
- Lodder WJ, Buisman AM, Rutjes SA, Heijne JC, Teunis PF, de Roda Husman AM. Feasibility of quantitative environmental surveillance in poliovirus eradication strategies. Appl Environ Microbiol. 2012;78:3800–5. DOIPubMedGoogle Scholar
- Hovi T, Shulman LM, van der Avoort H, Deshpande J, Roivainen M, De Gourville EM. Role of environmental poliovirus surveillance in global polio eradication and beyond. Epidemiol Infect. 2012;140:1–13. DOIPubMedGoogle Scholar
- Rutjes SA, Italiaander R, van den Berg HH, Lodder WJ, de Roda Husman AM. Isolation and detection of enterovirus RNA from large-volume water samples by using the NucliSens miniMAG system and real-time nucleic acid sequence-based amplification. Appl Environ Microbiol. 2005;71:3734–40. DOIPubMedGoogle Scholar
- Skraber S, Italiaander R, Lodder W, de Roda Husman AM. Noroviruses in archival samples. Emerg Infect Dis. 2005;11:489–91. DOIPubMedGoogle Scholar
- Lodder WJ, van den Berg HH, Rutjes SA, de Roda Husman AM. Presence of enteric viruses in source waters for drinking water production in the Netherlands. Appl Environ Microbiol. 2010;76:5965–71. DOIPubMedGoogle Scholar
- Katoh K, Toh H. Improved accuracy of multiple ncRNA alignment by incorporating structural information into a MAFFT-based framework. BMC Bioinformatics. 2008;9:212. DOIPubMedGoogle Scholar
- Drummond AJ, Rambaut A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol. 2007;7:214. DOIPubMedGoogle Scholar
- Strimmer K, Pybus OG. Exploring the demographic history of DNA sequences using the generalized skyline plot. Mol Biol Evol. 2001;18:2298–305. DOIPubMedGoogle Scholar
- Drummond AJ, Rambaut A, Shapiro B, Pybus OG. Bayesian coalescent inference of past population dynamics from molecular sequences. Mol Biol Evol. 2005;22:1185–92. DOIPubMedGoogle Scholar
- Verma H, Chitambar SD, Gopalkrishna V. Circulation of Aichi virus genotype B strains in children with acute gastroenteritis in India. Epidemiol Infect. 2011;139:1687–91. DOIPubMedGoogle Scholar
- Schijven JF, Teunis PF, Rutjes SA, Bouwknegt M, de Roda Husman AM. QMRAspot: a tool for quantitative microbial risk assessment from surface water to potable water. Water Res. 2011;45:5564–76. DOIPubMedGoogle Scholar
- Lukashev AN, Drexler JF, Belalov IS, Eschbach-Bludau M, Baumgarte S, Drosten C. Genetic variation and recombination in Aichi virus. J Gen Virol. 2012;93:1226–35. DOIPubMedGoogle Scholar
- Bibby K, Viau E, Peccia J. Viral metagenome analysis to guide human pathogen monitoring in environmental samples. Lett Appl Microbiol. 2011;52:386–92. DOIPubMedGoogle Scholar
- Cantalupo PG, Calgua B, Zhao G, Hundesa A, Wier AD, Katz JP, Raw sewage harbors diverse viral populations. mBio. 2011;2:pii:e00180-11.
- Chang JT, Chen YS, Chen BC, Chao D, Chang TH. Complete genome sequence of the first Aichi virus isolated in Taiwan. Genome Announc. 2013;1:e00107–12.PubMedGoogle Scholar
- Kapoor A, Simmonds P, Dubovi EJ, Qaisar N, Henriquez JA, Medina J, Characterization of a canine homolog of human Aichivirus. J Virol. 2011;85:11520–5. DOIPubMedGoogle Scholar
- Li L, Pesavento PA, Shan T, Leutenegger CM, Wang C, Delwart E. Viruses in diarrhoeic dogs include novel kobuviruses and sapoviruses. J Gen Virol. 2011;92:2534–41. DOIPubMedGoogle Scholar