Volume 7, Number 3—June 2001
Research
Outbreak of Human Monkeypox, Democratic Republic of Congo, 1996 to 1997
Figure 2
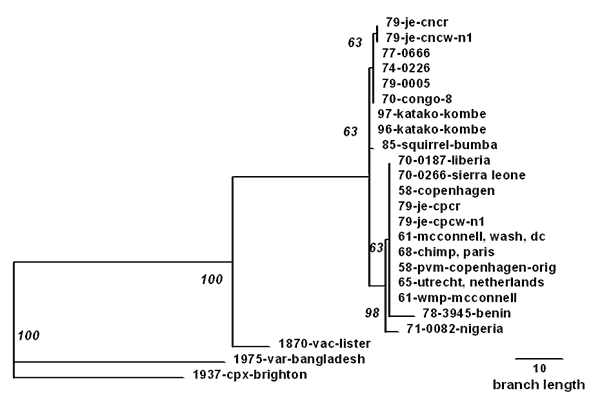
Figure 2. . Phylogenetic inference relationships of the open reading frames encoding the viral hemagglutinin protein of various monkeypox virus isolates and selected strains of vaccinia, variola, and cowpox viruses. Nucleotide sequences of polymerase chain reaction-generated amplicons were analyzed using PAUP parsimony analysis software version 3.1.1, as described (10). Parsimony analysis used 5,000 bootstraps and weighted the sequences for a transition-transversion ratio of 2 (bootstrap confidence intervals shown on branches).
References
- Marennikova SS, Seluhina EM, Mal'ceva NN, Cimiskjan KL, Macevic GR. Isolation and properties of the causal agent of a new variola-like disease (monkeypox) in man. Bull World Health Organ. 1972;46:599–611.PubMedGoogle Scholar
- Arita I, Jezek Z, Khodakevich L, Ruti K. Human monkeypox: a newly emerged orthopoxvirus zoonosis in the tropical rain forests of Africa. Am J Trop Med Hyg. 1985;34:781–9.PubMedGoogle Scholar
- Jezek Z, Fenner F. Human monkeypox. In: JL Melnick, editor. Monographs in virology. Volume 17. Basel: Karger; 1988.
- von Magnus P, Andersen EK, Petersen KB, Birch-Andersen A. A pox-like disease in cynomolgus monkeys. Acta Pathol Microbiol Scand. 1959;46:156–76. DOIGoogle Scholar
- Khodakevich L, Jezek Z, Messinger D. Monkeypox virus: ecology and public health significance. Bull World Health Organ. 1988;66:747–52.PubMedGoogle Scholar
- Khodakevich L, Szczeniowski M, Manbu-ma-Disu , Jezek Z, Marennikova S, Nakano J, The role of squirrels in sustaining monkeypox virus transmission. Trop Geogr Med. 1987;39:115–22.PubMedGoogle Scholar
- Fine PE, Jezek Z, Grab B, Dixon H. The transmission potential of monkeypox virus in human populations. Int J Epidemiol. 1988;17:643–50. DOIPubMedGoogle Scholar
- Jezek Z, Grab B, Szczeniowski MV, Paluku KM, Mutombo M. Human monkeypox: secondary attack rates. Bull World Health Organ. 1988;66:465–70.PubMedGoogle Scholar
- Jezek Z, Arita I, Mutombo M, Dunn C, Nakano JH, Szezeniowski M. Four generations of probable person-to-person transmission of human monkeypox. Am J Epidemiol. 1986;123:1004–12.PubMedGoogle Scholar
- Mukinda VB, Mwema G, Kilundu M, Heymann DL, Khan AS, Esposito JJ. Re-emergence of human monkeypox in Zaire in 1996. Lancet. 1997;349:1449–50. DOIPubMedGoogle Scholar
- Centers for Disease Control and Prevention. Human monkeypox- Zaire, 1996-1997. MMWR Morb Mortal Wkly Rep. 1997;46:304–7.PubMedGoogle Scholar
- Mills JN, Childs JE, Ksiazek TG, Peters CJ, Velleca WM. Methods for trapping and sampling small mammals for virologic testing. Atlanta: Centers for Disease Control and Prevention; 1995.
- Esposito JJ, Massung RF. Poxvirus infections in humans. In: Murray PR, Tenover F, Baron EJ, Pfaller MA, Yolken RH, editors. Manual of clinical microbiology. 6th ed. Washington: ASM Press;1995. p. 1131-8.
- Towbin H, Staehelin T, Gordon J. Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications. Proc Natl Acad Sci U S A. 1979;76:4350–4. DOIPubMedGoogle Scholar
- Loparev VN, Parsons JM, Knight JC, Panus JF, Ray CA, Buller RM, A third distinct tumor necrosis factor receptor of orthopoxviruses. Proc Natl Acad Sci U S A. 1998;95:3786–91. DOIPubMedGoogle Scholar
- Ropp SL, Esposito JJ, Loparev VN, Palumbo G. Poxviruses infecting humans. In: Murray PR, Barron CJ, Pfaller MA, Tenover FC, Yolken RH, editors. Manual of clinical microbiology. 7th ed. Washington: ASM Press; 1999. p. 1137-44.
- Ropp SL, Jin Q, Knight JC, Massung RF, Esposito JJ. PCR strategy for identification and differentiation of smallpox and other orthopoxviruses. J Clin Microbiol. 1995;33:2069–76.PubMedGoogle Scholar
- Esposito JJ, Knight JC. Orthopoxvirus DNA: a comparison of restriction profiles and maps. Virology. 1985;143:230–51. DOIPubMedGoogle Scholar
- Dean AG, Dean JA, Coulombier D, Burton AH, Brendel KA, Smith DC. Epi Info, version 6: A word-processing, database, and statistics program for public health on IBM-compatible microcomputers. Atlanta: Centers for Disease Control and Prevention; 1994.
- Fenner F, Wittek R, Dumbell KR. The Orthopoxviruses. San Diego: Academic Press; 1989. p. 162-5, 312-15.
- Ziegler DW, Hutchinson HD, Koplan JP, Nakano JH. Detection by radioimmunoassay of antibodies in human smallpox patients and vaccinees. J Clin Microbiol. 1975;1:311–7.PubMedGoogle Scholar
- Cherry JD, McIntosh K, Connor JD, Benenson AS, Alling DW, Rolfe UT, Primary percutaneous vaccination. J Infect Dis. 1977;135:145–54. DOIPubMedGoogle Scholar
- McIntosh K, Cherry JD, Benenson AS, Connor JD, Alling DW, Rolfe UT, Clinical and serologic study of four smallpox vaccines comparing variations of dose and route of administration. Standard percutaneous revaccination of children who receive primary percutaneous vaccination. J Infect Dis. 1977;135:155–66. DOIPubMedGoogle Scholar
- Jezek Z, Szczeniowski M, Paluku KM, Mutombo M, Grab B. Human monkeypox: confusion with chickenpox. Acta Trop. 1988;45:297–307.PubMedGoogle Scholar
- Centers for Disease Control and Prevention. Human monkeypox--Kasai Oriental, Democratic Republic of Congo, February 1996-October 1997. MMWR Morb Mortal Wkly Rep. 1997;46:1168–71.PubMedGoogle Scholar
Page created: April 26, 2012
Page updated: April 26, 2012
Page reviewed: April 26, 2012
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.