Volume 13, Number 6—June 2007
Dispatch
Reemergence of Oropouche Fever, Northern Brazil
Figure 2
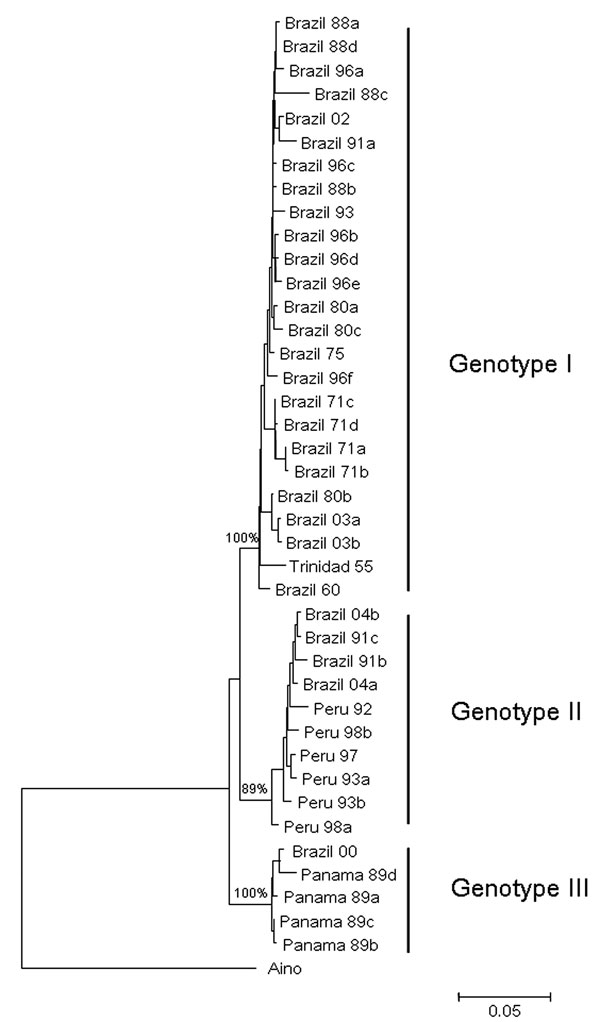
Figure 2. Comparative small (S) RNA phylogenetic tree constructed by using the neighbor-joining method for Oropouche virus strains isolated in Parauapebas and Porto de Moz, Pará State, Brazil. Bootstrap values were placed over the 3 nodes for each main group (I, II, and III). Aino virus S RNA sequence was used as an outgroup. Scale bar indicates a divergence of 5% in the nucleotide sequence.
Page created: June 30, 2010
Page updated: June 30, 2010
Page reviewed: June 30, 2010
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.