Volume 14, Number 9—September 2008
Research
Forest Fragmentation as Cause of Bacterial Transmission among Nonhuman Primates, Humans, and Livestock, Uganda
Figure 2
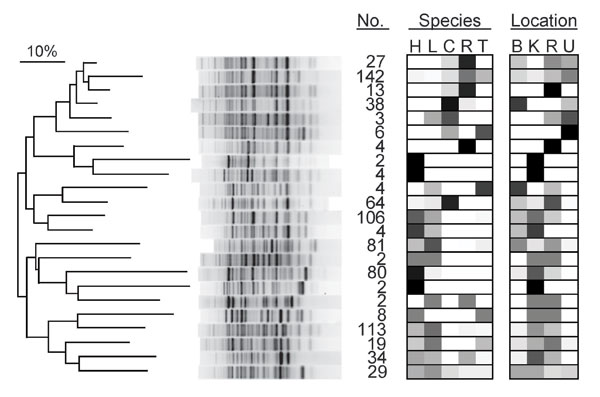
Figure 2. Dendrogram of genetic relatedness among 23 major clades of Escherichia coli from humans, domestic animals, and primates in 4 locations in and near Kibale National Park, western Uganda, derived from Rep-PCR genotypes. Major clades were identified from the full tree of 791 isolates by using the “cluster cutoff method” available in the computer program BioNumerics, version 4.0 (Applied Maths, Inc., Austin, TX, USA), which optimizes point-bisectional correlation across a range of cutoff similarity values to identify the most relevant clusters. A single representative bacterial genotype from each major clade is shown, and numbers of isolates falling within that clade are given (no.). Boxes indicate the host species and locations from which isolates in each clade were recovered and are shaded in proportion to the percentage of isolates in the clade from that species or location (0%, white; 100%, black). Species: H, human; L, livestock (cattle or goats); C, black-and-white colobus; R, red colobus; T, red-tailed guenon. Location: B, Bugembe fragment; K, Kiko 1 fragment; R, Rurama fragment; U, undisturbed locations within Kibale National Park. The dendrogram was drawn by using the neighbor-joining method (37) from a distance matrix generated from electrophoretic data that used optimized analytical parameters (30).
References
- Chapman CA, Gillespie TR, Goldberg TL. Primates and the ecology of their infectious diseases: how will anthropogenic change affect host-parasite interactions? Evol Anthropol. 2005;14:134–44. DOIGoogle Scholar
- Nunn CL, Altizer S. Infectious diseases in primates: behavior, ecology and evolution. Oxford: Oxford University Press; 2006.
- Leroy EM, Rouquet P, Formenty P, Souquiere S, Kilbourne A, Froment JM, Multiple Ebola virus transmission events and rapid decline of central African wildlife. Science. 2004;303:387–90. DOIPubMedGoogle Scholar
- Leendertz FH, Lankester F, Neel C, Drori O, Dupain J, Speede S, Anthrax joins Ebola in threatening central African great apes. Am J Primatol. 2006;68:928–33. DOIPubMedGoogle Scholar
- Kondgen S, Kuhl H, N’Goran PK, Walsh PD, Schenk S, Ernst N, Pandemic human viruses cause decline of endangered great apes. Curr Biol. 2008;18:260–4. DOIPubMedGoogle Scholar
- Leendertz FH, Pauli G, Maetz-Rensing K, Boardman W, Nunn C, Ellerbrok H, Pathogens as drivers of population declines: the importance of systematic monitoring in great apes and other threatened mammals. Biol Conserv. 2006;131:325–37. DOIGoogle Scholar
- Cowlishaw G, Dunbar R. Primate conservation biology. Chicago: University of Chicago Press; 2000.
- Chapman CA, Onderdonk DA. Forests without primates: primate/plant codependency. Am J Primatol. 1998;45:127–41. DOIPubMedGoogle Scholar
- Marsh LK, ed. Primates in fragments: ecology and conservation. New York: Kluwer Academic/Plenum Publishers; 2003.
- Estrada A, Coates-Estrada R. Tropical rain forest fragmentation and wild populations of primates at Los Tuxtlas, Mexico. Int J Primatol. 1996;17:759–83. DOIGoogle Scholar
- Onderdonk DA, Chapman CA. Coping with forest fragmentation: the primates of Kibale National Park, Uganda. Int J Primatol. 2000;21:587–611. DOIGoogle Scholar
- Gillespie TR, Chapman CA. Prediction of parasite infection dynamics in primate metapopulations based on attributes of forest fragmentation. Conserv Biol. 2006;20:441–8. DOIPubMedGoogle Scholar
- Gillespie TR, Chapman CA. Forest fragmentation, the decline of an endangered primate, and changes in host-parasite interactions relative to an unfragmented forest. Am J Primatol. 2008;70:222–30. DOIPubMedGoogle Scholar
- Salzer JS, Rwego IB, Goldberg TL, Kuhlenschmidt MS, Gillespie TR. Giardia sp. and Cryptosporidium sp. infections in primates in fragmented and undisturbed forest in western Uganda. J Parasitol. 2007;93:439–40. DOIPubMedGoogle Scholar
- Chapman CA, Wasserman MD, Gillespie TR, Speirs ML, Lawes MJ, Saj TL, Do food availability, parasitism, and stress have synergistic effects on red colobus populations living in forest fragments? Am J Phys Anthropol. 2006;131:525–34. DOIPubMedGoogle Scholar
- Donnenberg M, ed. Escherichia coli: virulence mechanisms of a versatile pathogen. San Diego: Academic Press; 2002.
- Beutin L. Emerging enterohaemorrhagic Escherichia coli, causes and effects of the rise of a human pathogen. J Vet Med B Infect Dis Vet Public Health. 2006;53:299–305. DOIPubMedGoogle Scholar
- Alekshun MN, Levy SB. Commensals upon us. Biochem Pharmacol. 2006;71:893–900. DOIPubMedGoogle Scholar
- Goldberg TL, Gillespie TR, Rwego IB, Wheeler E, Estoff EL, Chapman CA. Patterns of gastrointestinal bacterial exchange between chimpanzees and humans involved in research and tourism in western Uganda. Biol Conserv. 2007;135:511–7. DOIGoogle Scholar
- Struhsaker TT. Ecology of an African rain forest: logging in Kibale and the conflict between conservation and exploitation. Gainesville (FL): University Press of Florida; 1997.
- Chapman CA, Lambert JE. Habitat alteration and the conservation of African primates: case study of Kibale National Park, Uganda. Am J Primatol. 2000;50:169–85. DOIPubMedGoogle Scholar
- Chapman CA, Struhsaker TT, Lambert JE. Thirty years of research in Kibale National Park, Uganda, reveals a complex picture for conservation. Int J Primatol. 2005;26:539–55. DOIGoogle Scholar
- Struhsaker TT. Food habits of five monkey speices in the Kibale Forest, Uganda. In: Chivers DJ, Herbert J, editors. Recent advances in primatology. London: Academic Press; 1978. p. 225–48.
- Naughton-Treves L, Treves A, Chapman C, Wrangham R. Temporal patterns of crop-raiding by primates: linking food availability in croplands and adjacent forest. J Appl Ecol. 1998;35:596–606. DOIGoogle Scholar
- Feng P. Escherichia coli. In: Labbae RG, Garcia S, editors. Guide to foodborne pathogens. New York: John Wiley & Sons, Inc.; 2001. p. 143–62.
- Versalovic J, Koeuth T, Lupski JR. Distribution of repetitive DNA sequences in eubacteria and application to fingerprinting of bacterial genomes. Nucleic Acids Res. 1991;19:6823–31. DOIPubMedGoogle Scholar
- Johnson JR, O’Bryan TT. Improved repetitive-element PCR fingerprinting for resolving pathogenic and nonpathogenic phylogenetic groups within Escherichia coli. Clin Diagn Lab Immunol. 2000;7:265–73. DOIPubMedGoogle Scholar
- Carson CA, Shear BL, Ellersieck MR, Schnell JD. Comparison of ribotyping and repetitive extragenic palindromic-PCR for identification of fecal Escherichia coli from humans and animals. Appl Environ Microbiol. 2003;69:1836–9. DOIPubMedGoogle Scholar
- Goldberg TL, Gillespie TR, Singer RS. Optimization of analytical parameters for inferring relationships among Escherichia coli isolates from repetitive-element PCR by maximizing correspondence with multilocus sequence typing data. Appl Environ Microbiol. 2006;72:6049–52. DOIPubMedGoogle Scholar
- Excoffier L, Laval G, Schnieder S. Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evol Bioinform Online. 2005;1:47–50 [cited 2008 Jun 16]. Available from http://66.102.1.104/scholar?hl=en&lr=&q=cache:pIMXAyTZ-p8J:www.la-press.com/cr_data/files/f_EBO-1-ExcoffierSc_231.pdf+Arlequin+version+3.0
- Excoffier L, Smouse PE, Quattro JM. Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics. 1992;131:479–91.PubMedGoogle Scholar
- Reynolds J, Weir BS, Cockerham CC. Estimation of the coancestry coefficient: basis for a short-term genetic distance. Genetics. 1983;105:767–79.PubMedGoogle Scholar
- Felsenstein J. PHYLIP: Phylogenetic Inference Package, version 3.57c. Seattle: Department of Genetics, University of Washington; 1995.
- Maddison DR, Maddison WP. MacClade 4: Analysis of phylogeny and character evolution, version 4.0. Sunderland (MA): Sinauer Associates; 2000.
- Goldberg TL. Application of phylogeny reconstruction and character-evolution analysis to inferring patterns of directional microbial transmission. Prev Vet Med. 2003;61:59–70. DOIPubMedGoogle Scholar
- Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987;4:406–25.PubMedGoogle Scholar
- Goldberg TL, Gillespie TR, Rwego IB. Health and disease in the people, primates, and domestic animals of Kibale National Park: implications for conservation. In: Wrangham R, Ross E, editors. Science and conservation in African forests: the benefits of long-term research. Cambridge: Cambridge University Press; 2008. p. 75–87.
- Hartel PG, Summer JD, Segars WI. Deer diet affects ribotype diversity of Escherichia coli for bacterial source tracking. Water Res. 2003;37:3263–8. DOIPubMedGoogle Scholar
- Rivera SC, Hazen TC, Toranzos GA. Isolation of fecal coliforms from pristine sites in a tropical rain forest. Appl Environ Microbiol. 1988;54:513–7.PubMedGoogle Scholar