Volume 15, Number 10—October 2009
Letter
Aichi Virus Strains in Children with Gastroenteritis, China
Figure
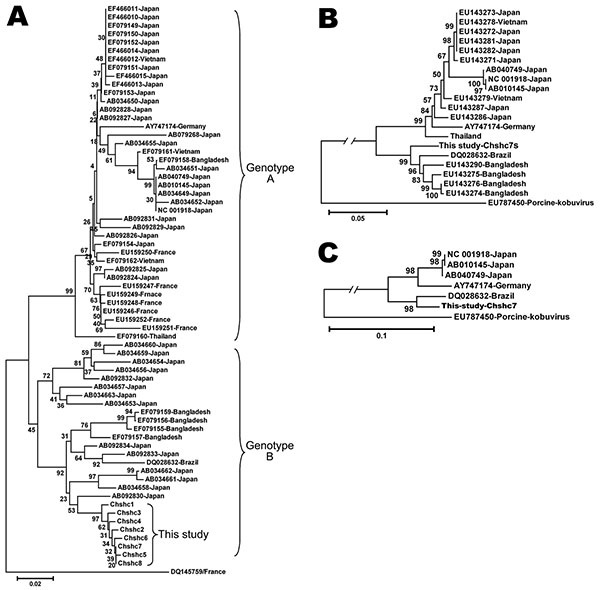
Figure. Phylogenetic tree constructed by using the neighbor-joining method and evaluated using the interior branch test method with MEGA4 software (www.megasoftware.net). Percentage of bootstrap support is indicated at each node. GenBank accession number, source, and country of origin are indicated. A) Phylogenetic tree constructed based on the 519-nt segment in the 3C/D junction region, a genotype C Aichi virus strain is included as an outgroup. Phylogenetic trees constructed from the capsid gene (B) and complete genome of Aichi virus (C); the porcine kobuvirus is included as an outgroup. Scale bars indicate nucleotide substitutions per site.
Page created: December 08, 2010
Page updated: December 08, 2010
Page reviewed: December 08, 2010
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.