Volume 15, Number 11—November 2009
Letter
Imported Chikungunya Virus Strains, Taiwan, 2006–2009
Figure
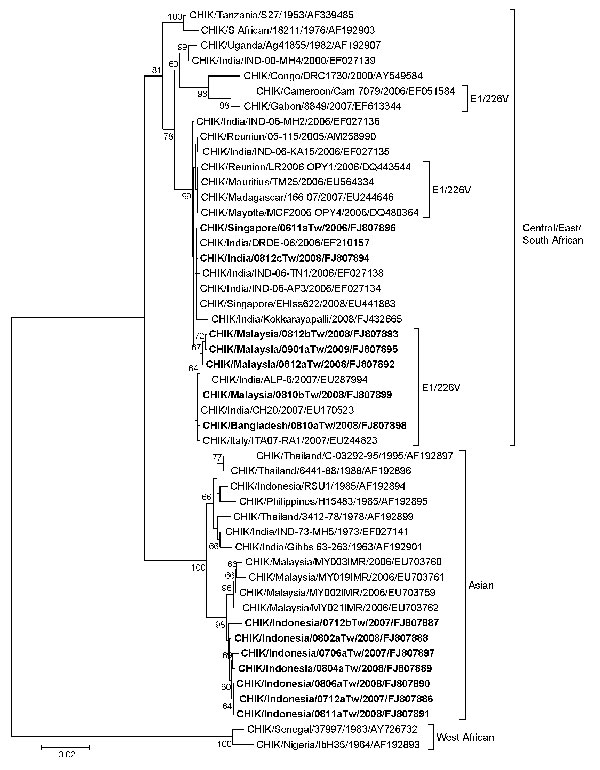
Figure. Phylogenetic relationships of chikungunya virus (CHIKV) isolates from 14 imported cases of chikungunya, Taiwan, 2006–2009. The tree was constructed on the basis of partial envelope 1 (E1) nucleotide sequences (836 bp, nt positions 10264–11099 of the prototype CHIKV S27 genomic sequence) of 49 CHIKV strains. Sequences obtained in this study are indicated in boldface. CHIKV strains with the E1-A226V mutation are indicated. Genotypes are indicated on the right. Viruses were identified by using the nomenclature of virus/country/strain/year of isolation/GenBank accession number. Analysis was performed by using MEGA 4 software and neighbor-joining (maximum composite likelihood) methods. Bootstrap support values >60 are shown (1,000 replicates). West African genotype Senegal strain 37997 sequence was used as the outgroup virus. Scale bar indicates nucleotide substitutions per site.