Novel Human Parechovirus from Brazil
Ana Maria Bispo de Filippis, Klaus Grywna, Andreas Stöcker, Patrícia Silva Almeida, Tereza Cristina Medrado Ribeiro, Monika Eschbach-Bludau, Nadine Petersen, Hugo da Costa Ribeiro, and Sung Sup Park

Author affiliations: University Hospital Prof Edgard Santos, Federal University of Bahia, Salvador, Brazil (J.F. Drexler, A. Stöcker); Bernhard Nocht Institute for Tropical Medicine, Hamburg, Germany (J.F. Drexler, N. Petersen); University of Bonn Medical Centre, Bonn, Germany (J.F. Drexler, K. Grywna, M. Eschbach-Bludau, C. Drosten); Federal University of Bahia, Salvador (P. Silva Almeida, T.C. Medrado Ribeiro, H. da Costa Ribeiro, Jr.)
Main Article
Figure 2
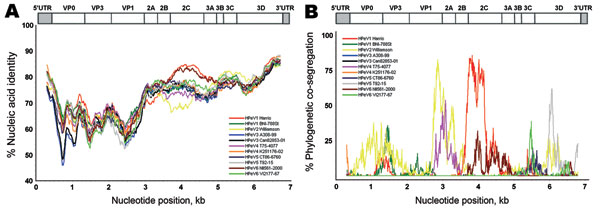
Figure 2. Nucleic acid identity with known parechoviruses. The near full-length genome of the new parechovirus BR/217/2006 was analyzed with SimPlot software (http://sray.med.som.jhmi.edu/SCRoftware/simplot) using a 600-bp sliding window and a step size of 10. Because of partially incomplete 5′ untranslated (UTR) region GenBank reference sequences, an approximate 400 nucleotides had to be cut from the 5′ end of all genomes. HPeV, human parechovirus. A) Nucleic acid identity, per analysis window, for strain BR/217/2006 with prototype strains. Nucleotide positions on x-axis show the center of the window. B) BootScan analysis using the same window settings. A bootstrapped phylogenetic analysis was conducted per window along the alignment. Graphs represent the percentage (bootstrap values) at which each strain cosegregates phylogenetically in the analysis window with strain BR/217/2006. Prototype strains used for comparison are shown in the inserts in each panel. A schematic representation of the parechovirus genome is given on top.
Main Article
Page created: December 08, 2010
Page updated: December 08, 2010
Page reviewed: December 08, 2010
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.
