Volume 15, Number 4—April 2009
Research
Exotic Small Mammals as Potential Reservoirs of Zoonotic Bartonella spp.
Figure
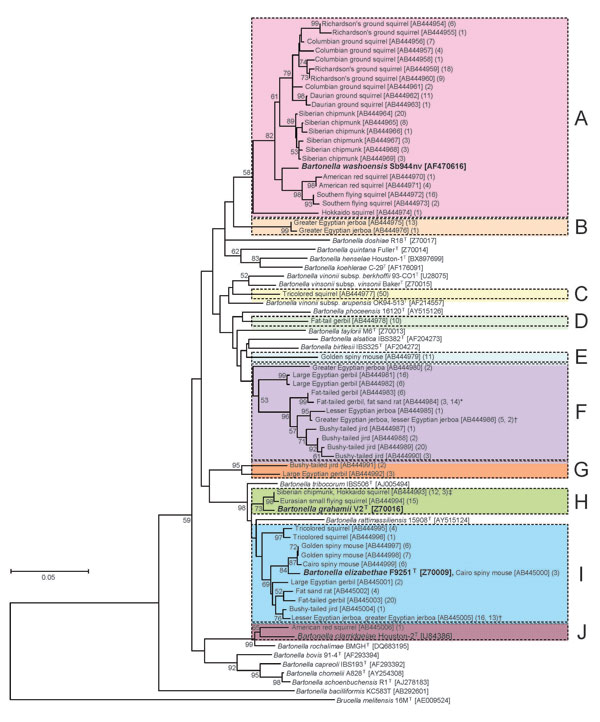
Figure. Phylogenetic tree based on a 312-bp region of the citrate synthase (gltA) gene sequence, constructed from Bartonella spp. isolates from 142 exotic small mammals imported into Japan as pets, June 2004–October 2007.Isolates from imported animals were compared with the type strains of known Bartonella spp. The phylogenetic tree was constructed by the neighbor-joining method, and bootstrap values were obtained with 1,000 replicates if values >50% were noted. The Brucella melitensis strain 16M sequence was used as an out-group. The GenBank accession number and the number of isolates are indicated in brackets and parentheses, respectively. The scale bar indicates 0.05 estimated nucleotide substitutions per site. Each colored column corresponds to genogroup A to J. Isolates showing identical genotypes were obtained from fat-tailed gerbils and fat sand rats (*), greater Egyptian jerboas and lesser Egyptian jerboas (†), and Siberian chipmunks and Hokkaido squirrels (‡).