Volume 15, Number 5—May 2009
Research
New Respiratory Enterovirus and Recombinant Rhinoviruses among Circulating Picornaviruses
Figure 1
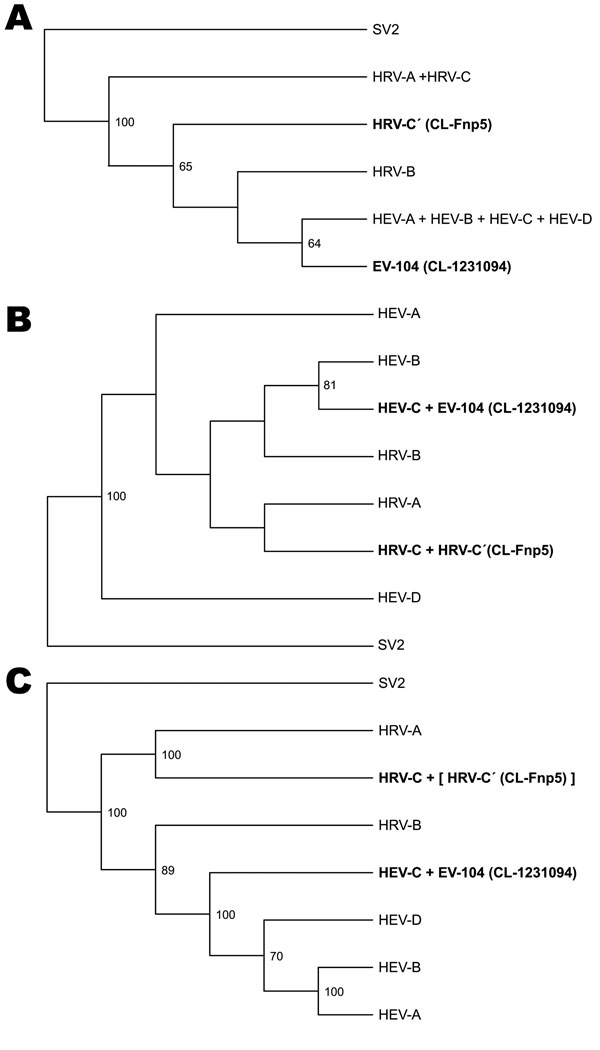
Figure 1. 5′ untranslated region (A), capsid protein VP1 (B), and complete genome (C) phylogeny of the virus clades studied. Trees were produced by condensing the full phylogeny shown in Technical Appendix 213) and a related clinical strain (CL-Fnp5). HRV-C includes the new clade described since 2006 (9–14,16). Enterovirus 104 (EV-104) and the related strain CL-1231094 refer to a previously unknown enterovirus clade described in this study. In panel C, HRV-C′ is shown in brackets to indicate its expected location (based on VP1 and 3D sequences). Simian picornavirus 1 (SV2) was used as an outgroup. HEV, human enterovirus. Bootstrap support values <50 are not shown in the trees. New viruses are shown in boldface.
References
- Tapparel C, Junier T, Gerlach D, Cordey S, Van Belle S, Perrin L, New complete genome sequences of human rhinoviruses shed light on their phylogeny and genomic features. BMC Genomics. 2007;8:224. DOIPubMedGoogle Scholar
- Newcombe NG, Andersson P, Johansson ES, Au GG, Lindberg AM, Barry RD, Cellular receptor interactions of C-cluster human group A coxsackieviruses. J Gen Virol. 2003;84:3041–50. DOIPubMedGoogle Scholar
- Pulli T, Koskimies P, Hyypia T. Molecular comparison of coxsackie A virus serotypes. Virology. 1995;212:30–8. DOIPubMedGoogle Scholar
- Dufresne AT, Gromeier M. A nonpolio enterovirus with respiratory tropism causes poliomyelitis in intercellular adhesion molecule 1 transgenic mice. Proc Natl Acad Sci U S A. 2004;101:13636–41. DOIPubMedGoogle Scholar
- Oberste MS, Maher K, Schnurr D, Flemister MR, Lovchik JC, Peters H, Enterovirus 68 is associated with respiratory illness and shares biological features with both the enteroviruses and the rhinoviruses. J Gen Virol. 2004;85:2577–84. DOIPubMedGoogle Scholar
- Ledford RM, Patel NR, Demenczuk TM, Watanyar A, Herbertz T, Collett MS, VP1 sequencing of all human rhinovirus serotypes: insights into genus phylogeny and susceptibility to antiviral capsid-binding compounds. J Virol. 2004;78:3663–74. DOIPubMedGoogle Scholar
- Laine P, Blomqvist S, Savolainen C, Andries K, Hovi T. Alignment of capsid protein VP1 sequences of all human rhinovirus prototype strains: conserved motifs and functional domains. J Gen Virol. 2006;87:129–38. DOIPubMedGoogle Scholar
- Savolainen C, Blomqvist S, Mulders MN, Hovi T. Genetic clustering of all 102 human rhinovirus prototype strains: serotype 87 is close to human enterovirus 70. J Gen Virol. 2002;83:333–40.PubMedGoogle Scholar
- Arden KE, McErlean P, Nissen MD, Sloots TP, Mackay IM. Frequent detection of human rhinoviruses, paramyxoviruses, coronaviruses, and bocavirus during acute respiratory tract infections. J Med Virol. 2006;78:1232–40. DOIPubMedGoogle Scholar
- Kistler A, Avila PC, Rouskin S, Wang D, Ward T, Yagi S, Pan-viral screening of respiratory tract infections in adults with and without asthma reveals unexpected human coronavirus and human rhinovirus diversity. J Infect Dis. 2007;196:817–25. DOIPubMedGoogle Scholar
- Lamson D, Renwick N, Kapoor V, Liu Z, Palacios G, Ju J, MassTag polymerase-chain-reaction detection of respiratory pathogens, including a new rhinovirus genotype, that caused influenza-like illness in New York State during 2004–2005. J Infect Dis. 2006;194:1398–402. DOIPubMedGoogle Scholar
- Lau SK, Yip CC, Tsoi HW, Lee RA, So LY, Lau YL, Clinical features and complete genome characterization of a distinct human rhinovirus (HRV) genetic cluster, probably representing a previously undetected HRV species, HRV-C, associated with acute respiratory illness in children. J Clin Microbiol. 2007;45:3655–64. DOIPubMedGoogle Scholar
- Lee WM, Kiesner C, Pappas T, Lee I, Grindle K, Jartti T, A diverse group of previously unrecognized human rhinoviruses are common causes of respiratory illnesses in infants. PLoS One. 2007;2:e966. DOIPubMedGoogle Scholar
- McErlean P, Shackelton LA, Lambert SB, Nissen MD, Sloots TP, Mackay IM. Characterisation of a newly identified human rhinovirus, HRV-QPM, discovered in infants with bronchiolitis. J Clin Virol. 2007;39:67–75. DOIPubMedGoogle Scholar
- McErlean P, Shackelton LA, Andrews E, Webster DR, Lambert SB, Nissen MD, Distinguishing molecular features and clinical characteristics of a putative new rhinovirus species, human rhinovirus C (HRV C). PLoS One. 2008;3:e1847.
- Renwick N, Schweiger B, Kapoor V, Liu Z, Villari J, Bullmann R, A recently identified rhinovirus genotype is associated with severe respiratory-tract infection in children in Germany. J Infect Dis. 2007;196:1754–60. DOIPubMedGoogle Scholar
- Cordey S, Gerlach D, Junier T, Zdobnov EM, Kaiser L, Tapparel C. The cis-acting replication elements define human enterovirus and rhinovirus species. RNA. 2008;14:1568–78. DOIPubMedGoogle Scholar
- Savolainen C, Laine P, Mulders MN, Hovi T. Sequence analysis of human rhinoviruses in the RNA-dependent RNA polymerase coding region reveals large within-species variation. J Gen Virol. 2004;85:2271–7. DOIPubMedGoogle Scholar
- Deffernez C, Wunderli W, Thomas Y, Yerly S, Perrin L, Kaiser L. Amplicon sequencing and improved detection of human rhinovirus in respiratory samples. J Clin Microbiol. 2004;42:3212–8. DOIPubMedGoogle Scholar
- Regamey N, Kaiser L, Roiha HL, Deffernez C, Kuehni CE, Latzin P, Viral etiology of acute respiratory infections with cough in infancy: a community-based birth cohort study. Pediatr Infect Dis J. 2008;27:100–5.PubMedGoogle Scholar
- Garbino J, Gerbase MW, Wunderli W, Deffernez C, Thomas Y, Rochat T, Lower respiratory viral illnesses: improved diagnosis by molecular methods and clinical impact. Am J Respir Crit Care Med. 2004;170:1197–203. DOIPubMedGoogle Scholar
- Kronenberg A, Zucs P, Droz S, Muhlemann K. Distribution and invasiveness of Streptococcus pneumoniae serotypes in Switzerland, a country with low antibiotic selection pressure, from 2001 to 2004. J Clin Microbiol. 2006;44:2032–8. DOIPubMedGoogle Scholar
- Allander T, Tammi MT, Eriksson M, Bjerkner A, Tiveljung-Lindell A, Andersson B. Cloning of a human parvovirus by molecular screening of respiratory tract samples. Proc Natl Acad Sci U S A. 2005;102:12891–6. DOIPubMedGoogle Scholar
- Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32:1792–7. DOIPubMedGoogle Scholar
- Rice P, Longden I, Bleasby A. EMBOSS: the European Molecular Biology Open Software Suite. Trends Genet. 2000;16:276–7. DOIPubMedGoogle Scholar
- Guindon S, Gascuel O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol. 2003;52:696–704. DOIPubMedGoogle Scholar
- Salminen MO, Carr JK, Burke DS, McCutchan FE. Identification of breakpoints in intergenotypic recombinants of HIV type 1 by bootscanning. AIDS Res Hum Retroviruses. 1995;11:1423–5. DOIPubMedGoogle Scholar
- Garbino J, Soccal PM, Aubert JD, Rochat T, Meylan P, Thomas Y, Respiratory viruses in bronchoalveolar lavage: a hospital-based cohort study in adults. Thorax. 2009; [Epub ahead of print].
- Brown B, Oberste MS, Maher K, Pallansch MA. Complete genomic sequencing shows that polioviruses and members of human enterovirus species C are closely related in the noncapsid coding region. J Virol. 2003;77:8973–84. DOIPubMedGoogle Scholar
- Nix WA, Oberste MS, Pallansch MA. Sensitive, seminested PCR amplification of VP1 sequences for direct identification of all enterovirus serotypes from original clinical specimens. J Clin Microbiol. 2006;44:2698–704. DOIPubMedGoogle Scholar
- Kistler AL, Webster DR, Rouskin S, Magrini V, Credle JJ, Schnurr DP, Genome-wide diversity and selective pressure in the human rhinovirus. Virol J. 2007;4:40. DOIPubMedGoogle Scholar
- Simmonds P. Recombination and selection in the evolution of picornaviruses and other mammalian positive-stranded RNA viruses. J Virol. 2006;80:11124–40. DOIPubMedGoogle Scholar
- Smura T, Blomqvist S, Paananen A, Vuorinen T, Sobotova Z, Bubovica V, Enterovirus surveillance reveals proposed new serotypes and provides new insight into enterovirus 5′-untranslated region evolution. J Gen Virol. 2007;88:2520–6. DOIPubMedGoogle Scholar
- Oberste MS, Maher K, Michele SM, Belliot G, Uddin M, Pallansch MA. Enteroviruses 76, 89, 90 and 91 represent a novel group within the species Human enterovirus A. J Gen Virol. 2005;86:445–51. DOIPubMedGoogle Scholar
- Junttila N, Leveque N, Kabue JP, Cartet G, Mushiya F, Muyembe-Tamfum JJ, New enteroviruses, EV-93 and EV-94, associated with acute flaccid paralysis in the Democratic Republic of the Congo. J Med Virol. 2007;79:393–400. DOIPubMedGoogle Scholar
- Norder H, Bjerregaard L, Magnius L, Lina B, Aymard M, Chomel JJ. Sequencing of ‘untypable’ enteroviruses reveals two new types, EV-77 and EV-78, within human enterovirus type B and substitutions in the BC loop of the VP1 protein for known types. J Gen Virol. 2003;84:827–36. DOIPubMedGoogle Scholar
- Witso E, Palacios G, Cinek O, Stene LC, Grinde B, Janowitz D, High prevalence of human enterovirus a infections in natural circulation of human enteroviruses. J Clin Microbiol. 2006;44:4095–100. DOIPubMedGoogle Scholar
- Jiang P, Faase JA, Toyoda H, Paul A, Wimmer E, Gorbalenya AE. Evidence for emergence of diverse polioviruses from C-cluster coxsackie A viruses and implications for global poliovirus eradication. Proc Natl Acad Sci U S A. 2007;104:9457–62. DOIPubMedGoogle Scholar
- Domingo E, Martin V, Perales C, Escarmis C. Coxsackieviruses and quasispecies theory: evolution of enteroviruses. Curr Top Microbiol Immunol. 2008;323:3–32. DOIPubMedGoogle Scholar
- Lukashev AN. Role of recombination in evolution of enteroviruses. Rev Med Virol. 2005;15:157–67. DOIPubMedGoogle Scholar
Page created: December 16, 2010
Page updated: December 16, 2010
Page reviewed: December 16, 2010
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.