Volume 16, Number 1—January 2010
Dispatch
Human Listeriosis Caused by Listeria ivanovii
Figure
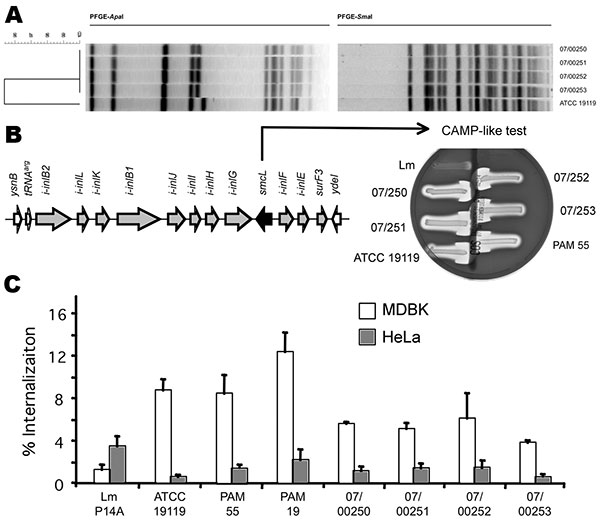
Figure. Characterization of the Listeria ivanovii subsp. ivanovii isolates from a 55-year-old man with gastroenteritis and bacteremia. A) The 4 isolates, 07/00250, 07/00251, and 07/00252 from blood, and 07/00253 from feces, were analyzed by pulsed-field gel electrophoresis (PFGE) with ApaI and SmaI restriction enzymes (9). The L. ivanovii subsp. ivanovii type strain American Type Culture Collection (ATCC) 19119 was used as control. Profiles were compared according to band positions by using the Dice coefficient and were clustered by using unweighted pair–group method averages. Criterion of dissimilarity = 1 band difference (maximum position tolerance 1.5%). B) L. ivanovii–specific virulence locus LIPI-2 and its phenotypic marker (sphingomyelinase production as shown by a CAMP-like test with an indicator strain of Rhodococcus equi on sheep blood agar). Left, genetic structure of LIPI-2. Arrowheads indicate positions of the oligonucleotide primers used in the 19 intragenic and intergenic PCRs to map the locus in the isolates; arrows represent genes (those belonging to LIPI-2 are gray, the sphingomyelinase gene is black, and flanking genes from the core listerial genome are white) (10). Right, typical shovel-shaped synergistic hemolysis reactions caused by L. ivanovii sphingomyelinase in the presence of R. equi cholesterol oxidase compared with the negative reaction given by L. monocytogenes (Lm). C) Invasion (gentamicin protection) assays in bovine (Madin-Darby bovine kidney) and human (HeLa) epithelial cells. The human isolates were compared with ruminant isolates ATCC 19119, PAM 55, and PAM 19 and with the L. monocytogenes strain P14A. Error bars indicate SEM of at least 2 duplicate experiments.
References
- Seeliger HPR, Jones D. Genus Listeria. In: Sneath PHA, Mair NS, Sharpe ME, and Holt JG, editors. Bergey’s manual of systematic bacteriology, Vol. 2. Baltimore: Williams & Wilkins; 1986. p. 1235–45.
- Vázquez-Boland JA, Kuhn M, Berche P, Chakraborty T, Domínguez-Bernal G, Goebel W, Listeria pathogenesis and molecular virulence determinants. Clin Microbiol Rev. 2001;14:584–640. DOIPubMedGoogle Scholar
- Busch LA. Human listeriosis in the United States, 1967–1969. J Infect Dis. 1971;123:328–32.PubMedGoogle Scholar
- Rocourt J, Seeliger HP. Distribution des espèces du genre Listeria. Zentralbl Bakteriol Mikrobiol Hyg [A]. 1985;259:317–30.PubMedGoogle Scholar
- Elischerova K, Cupkova E, Urgeova E, Lysy J, Sesevickova A. Isolation of Listeria ivanovii in Slovakia [in Slovak]. Cesk Epidemiol Mikrobiol Imunol. 1990;39:228–36.PubMedGoogle Scholar
- Lecuit M, Dramsi S, Gottardi C, Fedor-Chaiken M, Gumbiner B, Cossart P. A single amino acid in E-cadherin responsible for host specificity towards the human pathogen Listeria monocytogenes. EMBO J. 1999;18:3956–63. DOIPubMedGoogle Scholar
- Cummins AJ, Fielding AK, McLauchlin J. Listeria ivanovii infection in a patient with AIDS. J Infect. 1994;28:89–91. DOIPubMedGoogle Scholar
- Snapir YM, Vaisbein E, Nassar F. Low virulence but potentially fatal outcome—Listeria ivanovii. Eur J Intern Med. 2006;17:286–7. DOIPubMedGoogle Scholar
- Troxler R, von Graevenitz A, Funke G, Wiedemann B, Stock I. Natural antibiotic susceptibility of Listeria species: L. grayi, L. innocua, L. ivanovii, L. monocytogenes, L. seeligeri and L. welshimeri strains. Clin Microbiol Infect. 2000;6:525–35. DOIPubMedGoogle Scholar
- Lessing MP, Curtis GD, Bowler IC. Listeria ivanovii infection. J Infect. 1994;29:230–1. DOIPubMedGoogle Scholar
- Scortti M, Lacharme-Lora L, Wagner M, Chico-Calero I, Losito P, Vázquez-Boland JA. Coexpression of virulence and fosfomycin susceptibility in Listeria: molecular basis of an antimicrobial in vitro-in vivo paradox. Nat Med. 2006;12:515–7. DOIPubMedGoogle Scholar
- Ripio MT, Brehm K, Lara M, Suarez M, Vazquez-Boland JA. Glucose-1-phosphate utilization by Listeria monocytogenes is PrfA dependent and coordinately expressed with virulence factors. J Bacteriol. 1997;179:7174–80.PubMedGoogle Scholar
- Domínguez-Bernal G, Müller-Altrock S, González-Zorn B, Scortti M, Herrmann P, Monzó HJ, A spontaneous genomic deletion in Listeria ivanovii identifies LIPI-2, a species-specific pathogenicity island encoding sphingomyelinase and numerous internalins. Mol Microbiol. 2006;59:415–32. DOIPubMedGoogle Scholar
- Dalton CB, Austin CC, Sobel J, Hayes PS, Bibb WF, Graves LM, An outbreak of gastroenteritis and fever due to Listeria monocytogenes in milk. N Engl J Med. 1997;336:100–5. DOIPubMedGoogle Scholar
- Gaya P, Saralegui C, Medina M, Nuñez M. Occurrence of Listeria monocytogenes and other Listeria spp. in raw caprine milk. J Dairy Sci. 1996;79:1936–41.PubMedGoogle Scholar
1These authors contributed equally to this article.