Volume 17, Number 11—November 2011
Dispatch
Influenza B Viruses with Mutation in the Neuraminidase Active Site, North Carolina, USA, 2010–11
Figure
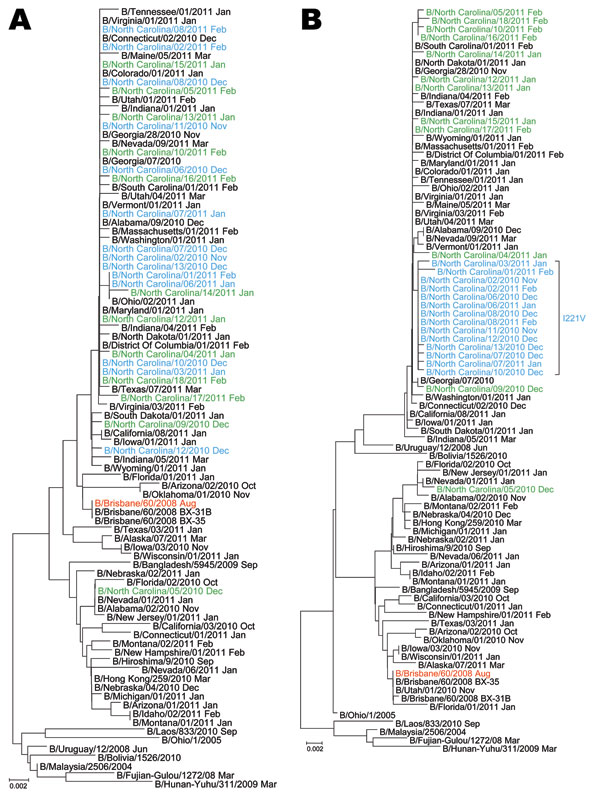
Figure. Phylogenetic analysis of A) hemagglutinin and B) neuraminidase genes of Victoria lineage type B influenza viruses (n = 89). Red indicates the 2010−2011 Northern Hemisphere vaccine strain; blue indicates the cluster of influenza B viruses identified in North Carolina carrying the I221V substitution in the neuraminidase; green indicates viruses collected from North Carolina with wild-type sequence at position 221 in the neuraminidase; black indicates representatives of globally circulating influenza B viruses. Month of collection is shown after virus strain designation. Evolutionary distances were computed by using the Tamura-Nei method (www.megasoftware.net/WebHelp/part_iv___evolutionary_analysis/computing_evolutionary_distances/distance_models/nucleotide_substitution_models/hc_tamura_nei_distance.htm). Scale bars indicate number of base substitutions per site.