Volume 17, Number 6—June 2011
CME ACTIVITY - Research
Cefepime-Resistant Pseudomonas aeruginosa
Cite This Article
Citation for Media
Abstract
Resistance to extended-spectrum cephalosporins complicates treatment of Pseudomonas aeruginosa infections. To elucidate risk factors for cefepime-resistant P. aeruginosa and determine its association with patient death, we conducted a case–control study in Philadelphia, Pennsylvania. Among 2,529 patients hospitalized during 2001–2006, a total of 213 (8.4%) had cefepime-resistant P. aeruginosa infection. Independent risk factors were prior use of an extended-spectrum cephalosphorin (p<0.001), prior use of an extended-spectrum penicillin (p = 0.005), prior use of a quinolone (p<0.001), and transfer from an outside facility (p = 0.01). Among those hospitalized at least 30 days, mortality rates were higher for those with cefepime-resistant than with cefepime-susceptible P. aeruginosa infection (20.2% vs. 13.2%, p = 0.007). Cefepime-resistant P. aeruginosa was an independent risk factor for death only for patients for whom it could be isolated from blood (p = 0.001). Strategies to counter its emergence should focus on optimizing use of antipseudomonal drugs.
1Current affiliation: Duke University School of Medicine, Durham, North Carolina, USA.
Pseudomonas aeruginosa is one of the most common gram-negative bacterial causes of health care–acquired infections (1–3). These infections result in high morbidity and mortality rates (4,5). When serious P. aeruginosa infections are suspected, early and appropriate antimicrobial drug therapy is crucial because inadequate drug selection has been associated with increased mortality rates (6,7). Complicating the empiric selection of adequate therapy is the increasing prevalence of antimicrobial drug resistance among P. aeruginosa (8–10). Even in initially susceptible strains, resistance can rapidly develop during treatment (11–13).
Cefepime, a fourth-generation cephalosphorin, is one of the few agents remaining that has reliable activity against P. aeruginosa. However, increased prevalence of resistance to cefepime among these organisms has been noted (14–18). As such, elucidating the epidemiology of cefepime-resistant P. aeruginosa is crucial to ensure that this agent remains a viable therapeutic option. Our goals were to identify risk factors for cefepime-resistant P. aeruginosa infections in the hospital setting and to describe the clinical effects of these infections.
The study was performed at the Hospital of the University of Pennsylvania (HUP), a 725-bed tertiary-care center, and Penn Presbyterian Medical Center (PPMC), a 344-bed urban community hospital. Each hospital is located in Philadelphia, Pennsylvania, USA, and is part of the University of Pennsylvania Health System. The study was reviewed and approved by the University of Pennsylvania Institutional Review Board.
Participants
To investigate risk factors for cefepime-resistant P. aeruginosa, we conducted a case–control study. We identified study participants through records obtained from the clinical microbiology laboratory at HUP, which performs bacterial cultures on all clinical specimens from HUP and PPMC. All adult patients with a positive P. aeruginosa culture result from January 1, 2001, through December 31, 2006, were eligible for inclusion. Each participant was included only one time; the first positive P. aeruginosa culture identified during the study period was used.
On the basis of our first study goal—identifying risk factors—we designated all participants with a cefepime-resistant P. aeruginosa–positive culture result as case-patients and all participants with a cefepime-susceptible P. aeruginosa culture result as controls. All eligible case-patients and controls were included according to the aforementioned eligibility criteria.
Variables
To assess risk factor variables, we used a comprehensive clinical and administrative University of Pennsylvania health system database, which contains data for all hospitalizations since January 1, 2001, and has been used successfully for similar studies of antimicrobial drug resistance (19–21). Data elements obtained were age, sex, race, hospital (HUP or PPMC), admission as a transfer from another facility (i.e., outside hospital, long-term care facility, rehabilitation center), location within the hospital at the time of culture (i.e., intensive care unit or not intensive care unit), length of hospital stay before culture, prior admission to HUP or PPMC within the past 30 days, Charlson index (22), and all-patient refined–diagnosis-related group (APR-DRG) classification. The following concurrent conditions were also noted: renal insufficiency (serum creatinine level >2.0 mg/dL or requirement for dialysis), malignancy, diabetes, cirrhosis, congestive heart failure, chronic pulmonary disease, immunosuppressive therapy, and HIV infection. These variables were based on International Classification of Diseases, Ninth Revision codes; laboratory data; and pharmacy data.
Drug Susceptibility Profiles
We documented antimicrobial drug susceptibility profiles, anatomic site of cultures, and any co-infections. Drug susceptibilities were conducted and interpreted by a semiautomated system (MicroScan WalkAway System, NC16 panel; Dade Behring, St. Louis, MO, USA) or disk-diffusion susceptibility testing in accordance with the criteria of the Clinical and Laboratory Standards Institute (23). Isolates with MIC = 16 (intermediate) or MIC >32 (resistant) were deemed resistant. A multidrug-resistant strain of P. aeruginosa was defined as a strain with resistance to >3 antimicrobial drug classes (24). We documented all antimicrobial drug treatment administered during the same inpatient admission for up to 30 days before the positive P. aeruginosa culture. We then categorized the drugs by individual agent, class, and spectrum of activity as follows: aminoglycosides (gentamicin, amikacin), quinolones (levofloxacin, ciprofloxacin), extended-spectrum penicillins (piperacillin-tazobactam), extended-spectrum cephalosporins (cefepime, ceftazidime), carbapenems (imipenem, meropenem), anaerobic therapy (amoxicillin/clavulanate, ampicillin/sulbactam, ceftriaxone, imipenem, meropenem, metronidazole, clindamycin), tetracyclines (doxycylcine), and macrolides (azithromycin, erythromycin) (25). During this study period, cefepime was the primary extended-spectrum cephalosporin used at HUP and PPMC, per formulary guidelines. For multivariable analyses, antimicrobial drugs were categorized by agent or class.
Mortality Rates
To assess the relationship between cefepime-resistant P. aeruginosa and mortality rates, we performed a retrospective cohort study, designating the participants with cefepime-resistant P. aeruginosa as the exposed group and those with cefepime-susceptible P. aeruginosa as the unexposed group. We focused specifically on rates for those hospitalized at least 30 days.
Statistical Analyses
We calculated the overall and annual prevalence of cefepime-resistant P. aeruginosa among all isolates identified during the study period. We then evaluated the annual prevalence of cefepime resistance over time by performing the χ2 test for trend (26).
To assess possible associations between potential risk factors and cefepime-resistant P. aeruginosa, we initially conducted bivariable analyses. Categorical variables were analyzed by using the Fisher exact test, and continuous variables were analyzed by using the Wilcoxon rank-sum test (27). The strength of each association was evaluated by calculating an odds ratio (OR) and a 95% confidence interval (CI). Multivariable analysis was performed by using forward stepwise multiple logistic regression (28). All variables with p<0.20 on bivariable analyses were considered for inclusion in the multivariable model. Backward stepwise multiple logistic regression was also performed to determine whether identification of risk factors varied with the approach to multivariable analysis. Because of the need to adjust for time at risk when investigating risk factors for antimicrobial drug resistance, we required the “duration of hospitalization prior to culture” variable to remain in the final model (29). We also analyzed the interaction between risk factor variables in the final model. Finally, to focus on those isolates likely to represent clinical infection, as per Centers for Disease Control and Prevention criteria, we repeated the analyses on blood isolates only (30).
To assess the association between cefepime-resistant P. aeruginosa and mortality rates for those hospitalized at least 30 days, we conducted bivariable and multivariable analyses in a similar fashion as for the case–control study. As we did for the case–control study, we repeated the analyses on blood isolates only.
We considered a 2-tailed p<0.05 significant. We used STATA version 10.0 (StataCorp, College Station, TX, USA) to perform the statistical analysis.
During the study period, culture results were positive for P. aeruginosa , and cefepime susceptibility was tested for 2,529 patients. Median patient age was 61 years (95% CI 60–62), and 1,439 (56.9%) patients were male. Regarding race and/or ethnicity, 1,116 (44.4%) were white, 848 (33.7%) were African American, 30 (1.2%) were Asian, 29 (1.2%) were Hispanic, and the rest were identified as other or unknown. Among all participants, 1,984 (78.5%) were hospitalized at HUP and 545 (21.6%) were hospitalized at PPMC.
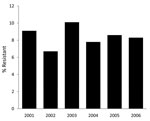
P. aeruginosa isolates came from the following anatomic sites: respiratory tract (247 [35.5%]), urine (763 [30.2%]), wound (467 [18.5%]), blood (248 [9.8%]), tissue (120 [4.7%]), and other (35 [1.3%]). Among the 2,529 isolates, 213 (8.4%) exhibited cefepime resistance and 339 (13.4%) exhibited multidrug resistance. Annual prevalence of P. aeruginosa cefepime resistance over time showed no significant trend (p = 0.99; Figure).
Using bivariate analysis to compare exposures, we found several differences between cefepime-resistant and cefepime-susceptible P. aeruginosa (Table 1). Specifically, participants with cefepime-resistant P. aeruginosa were more likely to have received an extended-spectrum cephalosporin, extended-spectrum penicillin, or quinolone. Multivariate analysis indicated that prior use of an extended-spectrum cephalosphorin had the strongest association with cefepime-resistant P. aeruginosa (adjusted OR 2.18, 95% CI 1.57–3.04; p<0.001) (Table 2). Independently associated with cefepime-resistant P. aeruginosa were prior use of an extended-spectrum penicillin or a quinolone and transfer from an outside facility (Table 2). No substantive differences were found in the final model when analyses were limited to blood isolates.
The overall mortality rate among participants was 13.8% (348/2,529). The mortality rate for participants with cefepime-resistant P. aeruginosa was 20.2% (43/213) and for participants with cefepime-susceptible P. aeruginosa was 13.2% (305/2,316) (relative risk [RR] 1.53, 95% CI 1.15– 2.04; p = 0.007). After controlling for significant confounders in the multivariate analysis, cefepime-resistant P. aeruginosa was no longer associated with death (Table 3). However, the association between cefepime-resistant P. aeruginosa and death varied significantly, depending on whether the isolate was from the blood or elsewhere. When analyses were restricted to blood isolates, a significant independent association between cefepime-resistant P. aeruginosa and death was found (RR 15.55, 95% CI 3.10–77.89; p = 0.001] (Table 4).
We found the following to be significant factors independently associated with isolation of a cefepime-resistant P. aeruginosa strain in culture of a clinical sample in the hospital setting: prior use of extended-spectrum cephalosphorins, extended-spectrum penicillins, or fluoroquinolones; and transfer from an outside facility. We also demonstrated that cefepime-resistant P. aeruginosa was independently associated with increased deaths among patients hospitalized for >30 days but only for those for whom P. aeruginosa was isolated from blood.
Past studies have found an association between use of an antipseudomonal agent and emergence of resistance to that same agent (19,31,32). Past studies have also demonstrated that P. aeruginosa resistance to 1 class of antimicrobial drugs is often associated with resistance to other classes (11,33). The tendency for health care–acquired P. aeruginosa to become resistant to drugs from multiple classes is well known, and several molecular mechanisms for its intrinsic and acquired resistance have been suggested (10). Our findings not only suggest that prior treatment with cefepime in itself is associated with subsequent emergence of cefepime-resistant P. aeruginosa but that even prior exposure to certain antipseudomonal agents in other classes is associated. Our results emphasize that to devise strategies that prevent further emergence of cefepime-resistant P. aeruginosa, recent prior use of antipseudomonal agents within the same class and from certain other classes must be recognized. The effect of curtailing use of non–β-lactam agents (i.e., fluoroquinolones) on cefepime-resistant P. aeruginosa prevalence should be formally assessed.
We also found that transfer from an outside facility was associated with cefepime-resistant P. aeruginosa infection. Because transferred patients potentially came from another hospital or from a long-term care facility, these patients might have been more likely to already be colonized with cefepime-resistant P. aeruginosa at the time of admission. That antimicrobial drug resistance is common in long-term care facilities is well known (34). Further work focusing specifically on antimicrobial drug–resistant P. aeruginosa infections in long-term care settings is warranted.
The significant association between cefepime-resistant P. aeruginosa infection and increased mortality rates (limited to those patients with P. aeruginosa bacteremia) may result from the fact that bacteremia is a more serious infection than, for example, a urinary tract infection. Alternatively, these results might be explained by noting that a blood isolate is more likely to represent a true infection than is an isolate from other anatomic sites, where isolates are more likely to represent colonization. Nonetheless, our results emphasize the potential serious effect of cefepime-resistant P. aeruginosa infection and the need for strategies to combat its further emergence.
This study has several potential limitations. The first is the ongoing, and appropriate, debate regarding the selection of the control group for case–control studies investigating the association between prior antimicrobial drug use and resistance. Like Harris and et al., we believe that selection of the control group depends on the study question (29,35). In our study, the main question was “What are the risk factors for cefepime resistance among all clinical isolates of P. aeruginosa in the hospital setting?” Thus, we selected patients with cefepime-susceptible P. aeruginosa infection as controls.
Another potential limitation is selection bias, which is always a concern in case–control studies. We believe that any such bias was minimized by the fact that every patient with a P. aeruginosa isolate was eligible for inclusion. Furthermore, all isolates were identified in the clinical microbiology laboratory at HUP, which processes all inpatient cultures at the participating study sites.
Misclassification bias is also a concern in case–control studies. However, the categorization of case-patients and controls and their exposure status was based entirely on preexisting clinical data from the clinical microbiology laboratory. The antimicrobial drug–susceptibility profiles were determined before study initiation, so determination of case and control status did not influence these profiles. Furthermore, case-patients and controls were selected without knowledge of their status regarding risk factors of interest. Thus, we believe any differential misclassification bias was unlikely.
Identification of participants in this study was based solely on clinical cultures. As such, that all of these cultures represented true infection is unlikely. For this reason, we performed additional analyses, focusing only on P. aeruginosa blood isolates because these would be expected to meet Centers for Disease Control and Prevention criteria for infection. The results of these secondary analyses did not differ substantively from the primary analyses investigating risk factors for cefepime-resistant P. aeruginosa. Finally, all patients in this study were admitted to either HUP or PPMC. Thus, our findings can only be generalized to similar academic centers. One must also keep in mind the differing resistance profiles at any given institution.
In conclusion, cefepime-resistant P. aeruginosa will negatively affect clinical outcomes, and strategies to counter its emergence are needed. Recognizing recent prior use of antipseudomonal agents, both within the same class and from certain other classes, is needed for devising successful interventions.
Dr Akhabue is a medical resident at Duke University, Durham, North Carolina, USA. His research interests focus on health care–acquired infections and antimicrobial drug resistance.
Acknowledgment
This study was supported by National Institutes of Health grant K24-AI080942 (to E.L.) and by a Commonwealth Universal Research Enhancement Program grant from the Pennsylvania State Department of Health (to E.L.).
References
- Gaynes R, Edwards JR. Overview of nosocomial infections caused by gram-negative bacilli. Clin Infect Dis. 2005;41:848–54. DOIPubMedGoogle Scholar
- Fluit AC, Jones ME, Schmitz FJ, Acar J, Gupta R, Verhoef J. Antimicrobial susceptibility and frequency of occurrence of clinical blood isolates in Europe from the SENTRY antimicrobial surveillance program, 1997 and 1998. Clin Infect Dis. 2000;30:454–60. DOIPubMedGoogle Scholar
- Streit JM, Jones RN, Sader HS, Fritsche TR. Assessment of pathogen occurrences and resistance profiles among infected patients in the intensive care unit: report from the SENTRY Antimicrobial Surveillance Program (North America, 2001). Int J Antimicrob Agents. 2004;24:111–8. DOIPubMedGoogle Scholar
- Crouch Brewer S, Wunderink RG, Jones CB, Leeper KV Jr. Ventilator-associated pneumonia due to Pseudomonas aeruginosa. Chest. 1996;109:1019–29. DOIPubMedGoogle Scholar
- Osmon S, Ward S, Fraser VJ, Kollef MH. Hospital mortality for patients with bacteremia due to Staphylococcus aureus or Pseudomonas aeruginosa. Chest. 2004;125:607–16. DOIPubMedGoogle Scholar
- Micek ST, Lloyd AE, Ritchie DJ, Reichley RM, Fraser VJ, Kollef MH. Pseudomonas aeruginosa bloodstream infection: importance of appropriate initial antimicrobial treatment. Antimicrob Agents Chemother. 2005;49:1306–11. DOIPubMedGoogle Scholar
- Vidal F, Mensa J, Almela M, Martínez JA, Marco F, Casals C, Epidemiology and outcome of Pseudomonas aeruginosa bacteremia, with special emphasis on the influence of antibiotic treatment. Analysis of 189 episodes. Arch Intern Med. 1996;156:2121–6. DOIPubMedGoogle Scholar
- National Nosocomial Infections Surveillance System. National Nosocomial Infections Surveillance (NNIS) System report, data summary from January 1992 through June 2004, issued October 2004. Am J Infect Control. 2004;32:470–85. DOIPubMedGoogle Scholar
- Karlowsky JA, Jones ME, Thornsberry C, Evangelista AT, Yee YC, Sahm DF. Stable antimicrobial susceptibility rates for clinical isolates of Pseudomonas aeruginosa from the 2001–2003 Tracking Resistance in the United States Today surveillance studies. Clin Infect Dis. 2005;40(Suppl 2):S89–98. DOIPubMedGoogle Scholar
- McGowan JE, Jr. Resistance in nonfermenting gram-negative bacteria: multidrug resistance to the maximum. Am J Med. 2006;119(6 Suppl 1):S29–36; discussion S62–70.
- Trouillet JL, Vuagnat A, Combes A, Kassis N, Chastre J, Gibert C. Pseudomonas aeruginosa ventilator-associated pneumonia: comparison of episodes due to piperacillin-resistant versus piperacillin-susceptible organisms. Clin Infect Dis. 2002;34:1047–54. DOIPubMedGoogle Scholar
- Carmeli Y, Troillet N, Eliopoulos GM, Samore MH. Emergence of antibiotic-resistant Pseudomonas aeruginosa: comparison of risks associated with different antipseudomonal agents. Antimicrob Agents Chemother. 1999;43:1379–82.PubMedGoogle Scholar
- Fink MP, Snydman DR, Niederman MS, Leeper KV Jr, Johnson RH, Heard SO, Treatment of severe pneumonia in hospitalized patients: results of a multicenter, randomized, double-blind trial comparing intravenous ciprofloxacin with imipenem-cilastatin. The Severe Pneumonia Study Group. Antimicrob Agents Chemother. 1994;38:547–57.PubMedGoogle Scholar
- American Thoracic Society; Infectious Diseases Society of America. Guidelines for the management of adults with hospital-acquired, ventilator-associated, and healthcare-associated pneumonia. Am J Respir Crit Care Med. 2005;171:388–416. DOIPubMedGoogle Scholar
- Neuhauser MM, Weinstein RA, Rydman R, Danziger LH, Karam G, Quinn JP. Antibiotic resistance among gram-negative bacilli in US intensive care units: implications for fluoroquinolone use. JAMA. 2003;289:885–8. DOIPubMedGoogle Scholar
- Ramphal R, Hoban DJ, Pfaller MA, Jones RN. Comparison of the activity of two broad-spectrum cephalosporins tested against 2,299 strains of Pseudomonas aeruginosa isolated at 38 North American medical centers participating in the SENTRY Antimicrobial Surveillance Program, 1997–1998. Diagn Microbiol Infect Dis. 2000;36:125–9. DOIPubMedGoogle Scholar
- Roberts JA, Webb SA, Lipman J. Cefepime versus ceftazidime: considerations for empirical use in critically ill patients. Int J Antimicrob Agents. 2007;29:117–28. DOIPubMedGoogle Scholar
- Sader HS, Fritsche TR, Jones RN. Potency and spectrum trends for cefepime tested against 65,746 clinical bacterial isolates collected in North American medical centers: results from the SENTRY Antimicrobial Surveillance Program (1998–2003). Diagn Microbiol Infect Dis. 2005;52:265–73. DOIPubMedGoogle Scholar
- Gasink LB, Fishman NO, Weiner MG, Nachamkin I, Bilker WB, Lautenbach E. Fluoroquinolone-resistant Pseudomonas aeruginosa: assessment of risk factors and clinical impact. Am J Med. 2006;119:526 e19–25.
- Lautenbach E, Synnestvedt M, Weiner MG, Bilker WB, Vo L, Schein J, Imipenem resistance in Pseudomonas aeruginosa: emergence, epidemiology, and impact on clinical and economic outcomes. Infect Control Hosp Epidemiol. 2010;31:47–53. DOIPubMedGoogle Scholar
- Lautenbach E, Weiner MG, Nachamkin I, Bilker WB, Sheridan A, Fishman NO. Imipenem resistance among Pseudomonas aeruginosa isolates: risk factors for infection and impact of resistance on clinical and economic outcomes. Infect Control Hosp Epidemiol. 2006;27:893–900. DOIPubMedGoogle Scholar
- Deyo RA, Cherkin DC, Ciol MA. Adapting a clinical comorbidity index for use with ICD-9-CM administrative databases. J Clin Epidemiol. 1992;45:613–9. DOIPubMedGoogle Scholar
- National Committee for Clinical Laboratory Standards (NCCLS). Performance standards for antimicrobial susceptibility testing. NCCLS approved standard M100–S11. Wayne (PA): The Committee; 2001.
- Falagas ME, Koletsi PK, Bliziotis IA. The diversity of definitions of multidrug-resistant (MDR) and pandrug-resistant (PDR) Acinetobacter baumannii and Pseudomonas aeruginosa. J Med Microbiol. 2006;55:1619–29. DOIPubMedGoogle Scholar
- MacAdam H, Zaoutis TE, Gasink LB, Bilker WB, Lautenbach E. Investigating the association between antibiotic use and antibiotic resistance: impact of different methods of categorising prior antibiotic use. Int J Antimicrob Agents. 2006;28:325–32. DOIPubMedGoogle Scholar
- Armitage P. Test for linear trend in proportions and frequencies. Biometrics. 1955;11:375–86. DOIGoogle Scholar
- Kleinbaum DK, Kupper LL, Morgenstern H. Epidemiologic research: principles and quantitative methods. New York: Van Nostrand Reinhold; 1982.
- Hosmer DLS. Applied logistic regression. New York: Wiley and Sons; 1989.
- Harris AD, Karchmer TB, Carmeli Y, Samore MH. Methodological principles of case–control studies that analyzed risk factors for antibiotic resistance: a systematic review. Clin Infect Dis. 2001;32:1055–61. DOIPubMedGoogle Scholar
- Garner JS, Jarvis WR, Emori TG, Horan TC, Hughes JM. CDC definitions for nosocomial infections, 1988. Am J Infect Control. 1988;16:128–40. DOIPubMedGoogle Scholar
- El Amari EB, Chamot E, Auckenthaler R, Pechère JC, Van Delden C. Influence of previous exposure to antibiotic therapy on the susceptibility pattern of Pseudomonas aeruginosa bacteremic isolates. Clin Infect Dis. 2001;33:1859–64. DOIPubMedGoogle Scholar
- Harris AD, Smith D, Johnson JA, Bradham DD, Roghmann MC. Risk factors for imipenem-resistant Pseudomonas aeruginosa among hospitalized patients. Clin Infect Dis. 2002;34:340–5. DOIPubMedGoogle Scholar
- López-Dupla M, Martínez JA, Vidal F, Almela M, Soriano A, Marco F, Previous ciprofloxacin exposure is associated with resistance to β-lactam antibiotics in subsequent Pseudomonas aeruginosa bacteremic isolates. Am J Infect Control. 2009;37:753–8. DOIPubMedGoogle Scholar
- Strausbaugh LJ, Crossley KB, Nurse BA, Thrupp LD. Antimicrobial resistance in long-term-care facilities. Infect Control Hosp Epidemiol. 1996;17:129–40. DOIPubMedGoogle Scholar
- Harris AD, Samore MH, Lipsitch M, Kaye KS, Perencevich E, Carmeli Y. Control-group selection importance in studies of antimicrobial resistance: examples applied to Pseudomonas aeruginosa, enterococci, and Escherichia coli. Clin Infect Dis. 2002;34:1558–63. DOIPubMedGoogle Scholar
Figure
Tables
Follow Up
Earning Medscape CME Credit
To obtain credit, you should first read the journal article. After reading the article, you should be able to answer the following, related, multiple-choice questions. To complete the questions and earn continuing medical education (CME) credit, please go to www.medscape.org/journal/eid. Credit cannot be obtained for tests completed on paper, although you may use the worksheet below to keep a record of your answers. You must be a registered user on Medscape.org. If you are not registered on Medscape.org, please click on the New Users: Free Registration link on the left hand side of the website to register. Only one answer is correct for each question. Once you successfully answer all post-test questions you will be able to view and/or print your certificate. For questions regarding the content of this activity, contact the accredited provider, CME@medscape.net. For technical assistance, contact CME@webmd.net. American Medical Association's Physician's Recognition Award (AMA PRA) credits are accepted in the US as evidence of participation in CME activities. For further information on this award, please refer to http://www.ama-assn.org/ama/pub/category/2922.html. The AMA has determined that physicians not licensed in the US who participate in this CME activity are eligible for AMA PRA Category 1 Credits™. Through agreements that the AMA has made with agencies in some countries, AMA PRA credit is acceptable as evidence of participation in CME activities. If you are not licensed in the US and want to obtain an AMA PRA CME credit, please complete the questions online, print the certificate and present it to your national medical association.
Cefepime-Resistant Pseudomonas aeruginosa
Medscape CME Questions
1. You are seeing a 61-year-old man admitted for pneumonia. His blood culture is now growing Pseudomonas aeruginosa, and you are concerned regarding the possibility of antimicrobial resistance of this organism.
What was the approximate rate of resistance to cefepime among isolates of P. aeruginosa in the current study?
A. Less than 1%
B. 8%
C. 22%
D. 47%
2. Which of the following variables independently increased the risk for P. aeruginosa resistance to cefepime in the current study?
A. Male sex
B. Diagnosis of pneumonia
C. Higher Charlson index score
D. Transfer from another facility
3. The patient was treated with antibiotics as an outpatient prior to hospital admission. Prior treatment with which classes of antibiotics was found to increase the risk for cefepime-resistant P. aeruginosa (CRPA) in the current study?
A. Aminoglycosides only
B. Extended-spectrum cephalosporins only
C. Extended-spectrum cephalosporins, extended-spectrum penicillins, and quinolones
D. Aminoglycosides, extended-spectrum penicillins, and macrolides
4. The patient is diagnosed with CRPA. What does the current study suggest regarding the effect of CRPA vs. cefepime-sensitive P. aeruginosa on the risk for mortality?
A. CRPA did not confer a higher risk for mortality in any analysis
B. Only older patients with CRPA were at a higher risk for death
C. Only patients with blood isolates for CRPA were at a higher risk for death
D. Any infection with CRPA was associated with a higher risk for death
Activity Evaluation
| 1. The activity supported the learning objectives. | ||||
| Strongly Disagree |
Strongly Agree
|
|||
|
1
|
2
|
3
|
4
|
5
|
| 2. The material was organized clearly for learning to occur. | ||||
| Strongly Disagree |
Strongly Agree
|
|||
|
1
|
2
|
3
|
4
|
5
|
| 3. The content learned from this activity will impact my practice. | ||||
| Strongly Disagree |
Strongly Agree
|
|||
|
1
|
2
|
3
|
4
|
5
|
| 4. The activity was presented objectively and free of commercial bias. | ||||
| Strongly Disagree |
Strongly Agree
|
|||
|
1
|
2
|
3
|
4
|
5
|
Related Links
Table of Contents – Volume 17, Number 6—June 2011
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Ebbing Lautenbach, University of Pennsylvania School of Medicine, Center for Clinical Epidemiology and Biostatistics, 825 Blockley Hall, 423 Guardian Dr, Philadelphia, PA 19104-6021, USA
Top