Volume 19, Number 4—April 2013
Research
Predicting Hotspots for Influenza Virus Reassortment
Figure 3
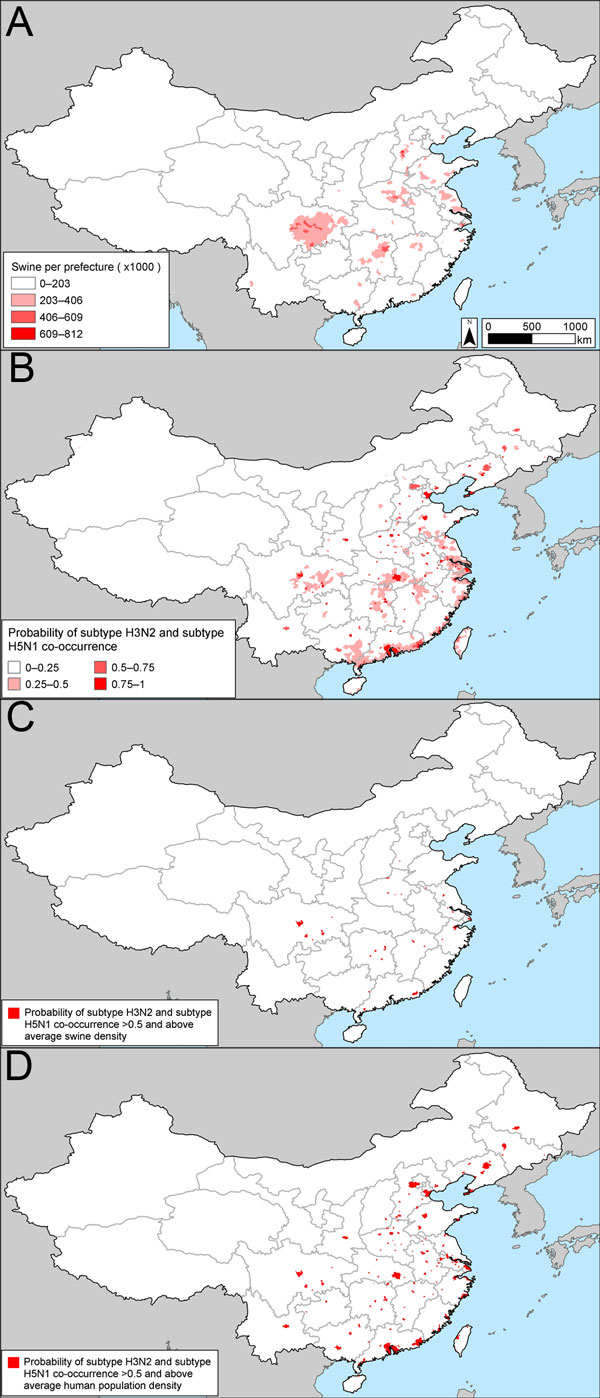
Figure 3. . . Potential influenza reassortment areas in People’s Republic of China determined by using the influenza virus subtype H5N1 outbreak dataset. A) Density of swine. B) Spatial model of the risk for subtype H3N2 and H5N1 co-occurrence according to the outbreak dataset. C) Areas with a probability of subtype H5N1 and H3N2 co-occurrence >50% and above average swine density. D) Areas with a probability of subtype H5N1 and H3N2 co-occurrence >50% and above average human population density. See , for corresponding maps based on the subtype H5N1 surveillance dataset.
Page created: March 13, 2013
Page updated: March 13, 2013
Page reviewed: March 13, 2013
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.