Volume 19, Number 6—June 2013
Letter
Crimean-Congo Hemorrhagic Fever Asia-2 Genotype, Pakistan
Figure
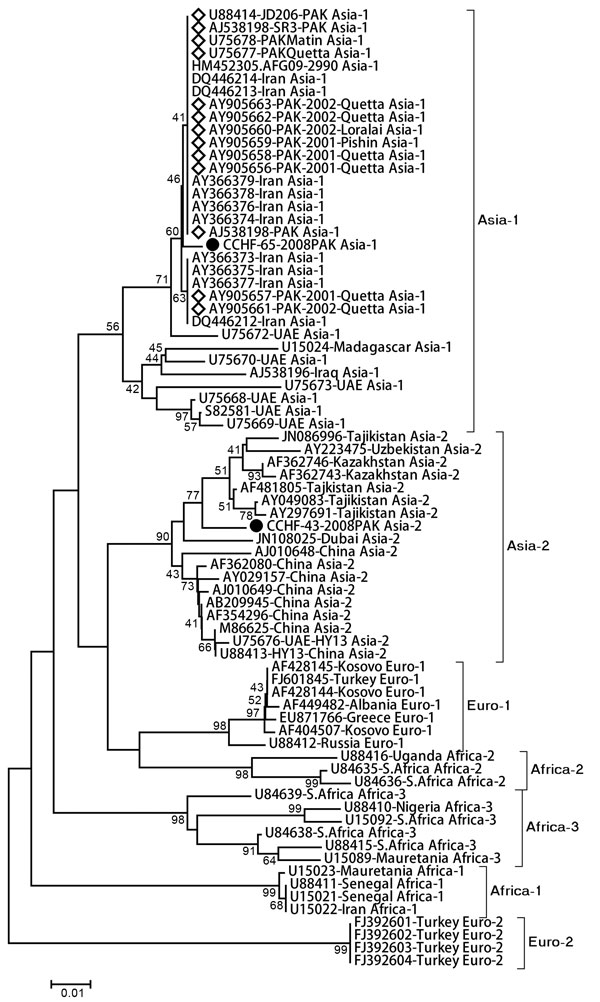
Figure. . . Phylogenetic analysis of partial small gene fragment (220 bp) obtained from Crimean-Congo hemorrhagic fever virus strains analyzed in this study (black circles). Reference strains belong to different genogroups as retrieved from GenBank. Diamonds indicate virus sequences previously reported from Pakistan. Evolutionary tree and distances (scale bar indicates number of base substitutions per site) were generated with the maximum composite likelihood method with Kimura-2 parameter distances by using MEGA 4.0 (http://megasoftware.net/). Numbers next to branches indicate the percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (1,000 replicates). The GenBank accession numbers, country, year of sample collection, and respective genotype information have been provided where available.