Volume 19, Number 9—September 2013
Research
Plasmodium falciparum Mutant Haplotype Infection during Pregnancy Associated with Reduced Birthweight, Tanzania
Figure 1
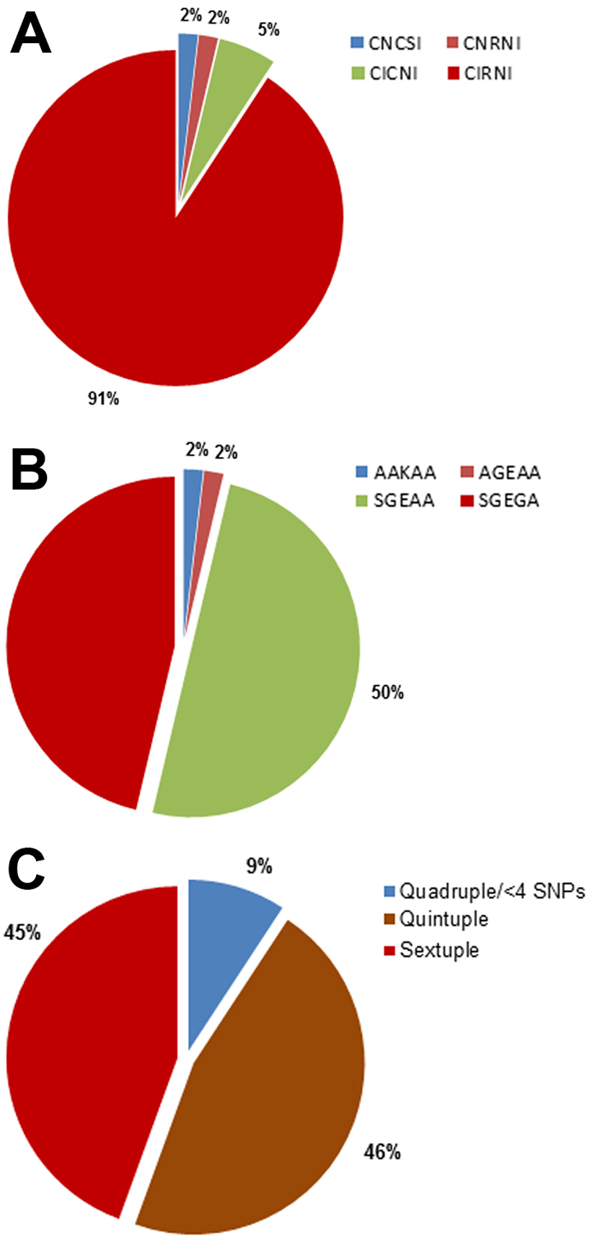
Figure 1. . . . Pfdhfr/Pfdhps haplotyping results for Plasmodium falciparum parasite isolates from 54 parasite isolates in 49 pregnant women, Korogwe District, Tanga Region, Tanzania, September 2008–October 2010. A) Proportion of single nucleotide polymorphisms (SNPs) conferring sulfadoxine–pyrimethamine resistance on the P. falciparum dihydrofolate reductase gene (Pfdhfr) at codons C50, N51I, C59R, S108N, and L164I, resulting in allelic haplotypes CNCSI (wild-type), CICNI (double Pfdhfr mutant), CNRNI (double Pfdhfr mutant), and CIRNI (triple Pfdhfr mutant). B) Proportion of SNPs conferring sulfadoxine–pyrimethamine resistance on the P. falciparum dihydropteroate synthetase (Pfdhps) gene at codons S436A, A437G, K540E, A613S/T, and A581G with allelic haplotypes AKAA (wild-type), AGEAA (double Pfdhps mutant), SGEAA (double Pfdhps mutant), and SGEGA (triple Pfdhps mutant). C) Proportions of Pfdhfr/Pfdhps quadruple or less, quintuple, and sextuple mutant haplotypes from the cohort of pregnant women. The derivations of the allelic haplotypes were based on a combination of 2 or 3 Pfdhfr SNPs with 2 or 3 Pfdhps SNPs forming quadruple (4 SNPs), quintuple (5 SNPs), and sextuple (6 SNPs) haplotypes. Quadruple haplotype or less included 4 SNPs (quadruple) and triple Pfdhfr (CIRNI, n = 1) or double dhps (SGEAA, n = 1) with wild-type dhfr (CNCSI, n = 1) or wild-type dhps (AKAA, n = 1). Less than 4 haplotypes had 1 triple or double mutation in 1 gene combined with a wild-type mutation in the other gene.
1Current affiliation: Mère et Enfant Face aux Infections Tropicales, Paris, France.