Real-Time Whole-Genome Sequencing for Surveillance of Listeria monocytogenes, France
Alexandra Moura
1, Mathieu Tourdjman
1, Alexandre Leclercq
1, Estelle Hamelin, Edith Laurent, Nathalie Fredriksen, Dieter Van Cauteren, Hélène Bracq-Dieye, Pierre Thouvenot, Guillaume Vales, Nathalie Tessaud-Rita, Mylène M. Maury, Andreea Alexandru, Alexis Criscuolo, Emmanuel Quevillon, Marie-Pierre Donguy, Vincent Enouf, Henriette de Valk, Sylvain Brisse
2, and Marc Lecuit
2
Author affiliations: Institut Pasteur, Paris, France (A. Moura, A. Leclercq, H. Bracq-Dieye, P. Thouvenot, G. Vales, N. Tessaud-Rita, M. Maury, A. Alexandru, A. Criscuolo, E. Quevillon, V. Enouf, S. Brisse, M. Lecuit); Institut National de la Santé et de la Recherche Médicale Unité 1117, Paris (A. Moura, M.M. Maury, M. Lecuit); Santé Publique France, Saint-Maurice, France (M. Tourdjman, E. Laurent, D. Van Cauteren, H. de Valk); Ministry of Agriculture, Agrifood, and Forestry, Paris (E. Hamelin, N. Fredriksen, M.-P. Donguy); Paris Descartes University, Paris (M. Lecuit); Necker-Enfants Malades University Hospital, Paris (M. Lecuit)
Main Article
Figure 1
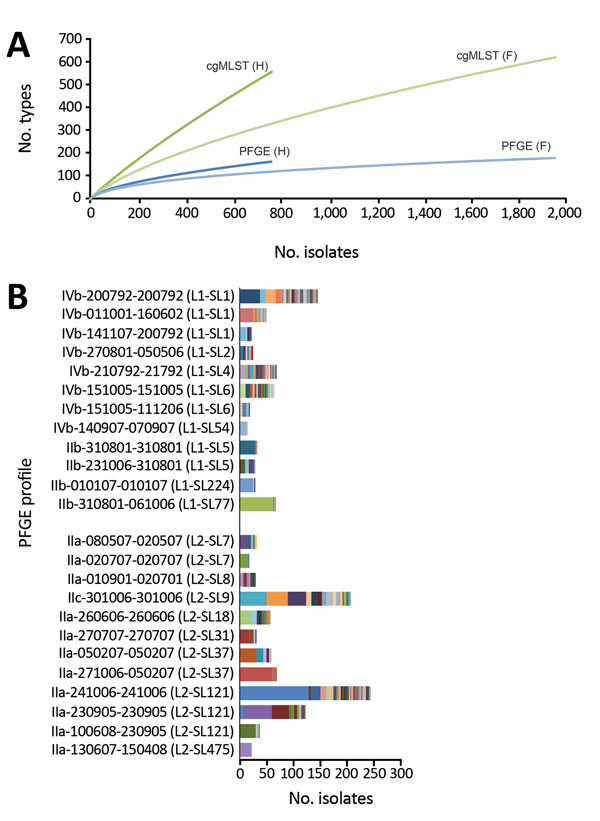
Figure 1. Discriminatory power of pulsed-field gel electrophoresis (PFGE) and core genome multilocus sequence typing (cgMLST) for surveillance of Listeria monocytogenes, France. A) Rarefaction analysis of type richness within human (H) and food-associated (F) isolates based on PFGE and cgMLST typing. B) Distribution of number of isolates per PFGE type and cgMLST subtyping. Only the most prevalent PFGE profiles (>20 isolates) are shown. Within each PFGE type, different cgMLST types (CTs) are represented by different arbitrary colors. PFGE types are coded by using the National Reference Centre for Listeria internal nomenclature of AscI-ApaI combined profiles. Information on lineage (L) and sublineage (SL) defined by cgMLST is provided for each PFGE type in parentheses.
Main Article
Page created: August 14, 2017
Page updated: August 14, 2017
Page reviewed: August 14, 2017
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.
