Volume 8, Number 12—December 2002
Dispatch
Rat-to-Human Transmission of Cowpox Infection
Figure 2
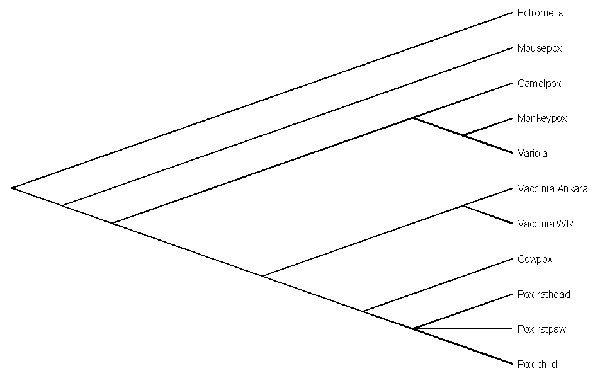
Figure 2. Phylogenetic tree of nucleotide sequences of 163-bp Orthopoxvirus fusion gene amplicons from the patient and rat (head and paw), Cowpox virus, Vaccinia virus (strain Ankara and WR), Camelpox virus, Monkeypox virus, Variola virus, and Ectromelia virus (Mousepox virus). The nucleotide sequences were aligned by using BioEdit software package (T. Hall, Dept. of Microbiology, Raleigh, NC). Phylogenetic relationships were determined by using the Lasergene software packages (DNASTAR Inc., Madison, WI).
Page created: July 19, 2010
Page updated: July 19, 2010
Page reviewed: July 19, 2010
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.