Volume 19, Number 7—July 2013
Research
Mutation in Spike Protein Cleavage Site and Pathogenesis of Feline Coronavirus
Figure 2
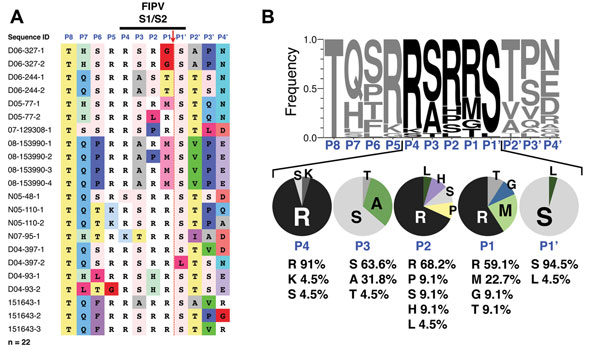
Figure 2. . Sequence analysis of feline infectious peritonitis virus (FIPV) spike S1/S2 site. RNA from 22 FIPVs collected from 11 cats who had feline infectious peritonitis was extracted, purified, and reverse-transcribed into cDNA. Sequencing of the spike gene was performed in a region surrounding the S1/S2 cleavage site. A) Sequence alignment. Sequence identification row (blue font): residue positions in the S1/S2 cleavage site from P8 to P4′. Red arrow indicates the site of furin cleavage. B) To visualize the diversity of residues at each position of the S1/S2 site, sequences were subjected to WebLogo 3.1 analysis (http://weblogo.threeplusone.com/create.cgi). Top: WebLogo for the 22 FIPV S1/S2 sequences with the frequency of residue found at each position displayed. Bottom: summary of the diversity of residues for each position from P4 to P1′ and percentages of each amino acid represented.
1These authors contributed equally to this article.