Volume 10, Number 10—October 2004
Perspective
Strain Typing Methods and Molecular Epidemiology of Pneumocystis Pneumonia
Abstract
Pneumocystis pneumonia (PCP) caused by the opportunistic fungal agent Pneumocystis jirovecii (formerly Pneumocystis carinii) continues to cause illness and death in HIV-infected patients. In the absence of a culture system to isolate and maintain live organisms, efforts to type and characterize the organism have relied on polymerase chain reaction–based approaches. Studies using these methods have improved understanding of PCP epidemiology, shedding light on sources of infection, transmission patterns, and potential emergence of antimicrobial resistance. One concern, however, is the lack of guidance regarding the appropriateness of different methods and standardization of these methods, which would facilitate comparing results reported by different laboratories.
Pneumocystis pneumonia (PCP) has been known for many years to be a disease of immunocompromised persons. Before the AIDS epidemic, it had been reported as a cause of death in malnourished infants (1). No standardized in vitro propagation system is currently available; consequently, much of the basic biology and epidemiology of Pneumocystis spp. remains poorly understood. Advances made over the last 15 years have been largely due to the use of molecular biologic approaches.
For almost 80 years, Pneumocystis jirovecii (formerly carinii) was considered to be a protozoan. In 1988, DNA studies clearly demonstrated that it was not a single species but a complex group of eukaryotic microorganisms, which were assigned to the kingdom Fungi (2–4) at the branch point between Ascomycota and Basidiomycota (5).
Many genetic typing methods use DNA sequencing approaches, but others use specific gene probes, single-strand conformation polymorphism (SSCP), or restriction fragment length polymorphisms (RFLP). Genetic typing has shown Pneumocystis biodiversity (6), environmental reservoirs (7,8), person-to-person transmission (9,10), recurrent infections (11), subclinical colonization and carriage (12–14), clinical manifestations (15), and sulfa exposure and suspected treatment or prophylaxis failures (16–19). These studies changed our epidemiologic understanding of PCP, and more studies now suggest collectively that a number of clinical PCP cases are newly acquired rather than activated latent infections (9–12,18).
In laboratories around the world, a number of typing procedures, each with its own strengths and weaknesses, are in use to address the clinical and epidemiologic issues discussed above. In the following sections, we discuss the most common methods, along with examples of how they have been used in molecular epidemiologic studies of PCP.
A variety of typing methods have been used for Pneumocystis genetic analysis, and a large number of gene loci have been examined. We focus on the methods and genes that have been most widely used for molecular epidemiologic analyses or have the greatest potential application.
Direct DNA sequence analysis is the most common approach currently used for Pneumocystis biodiversity and molecular typing studies. Sequence analysis of the thymidylate synthase (TS) and superoxide dismutase (SODA) gene loci, the EPSP synthase domain of the multifunctional arom gene, and the mitochondrial small subunit ribosomal RNA (mt SSU rRNA) locus have been used to distinguish Pneumocystis species from diverse mammalian hosts (6,20,21). Because of the generally low sequence divergence among P. jirovecii isolates at these loci, they are not highly discriminative for P. jirovecii typing. Several additional loci, however, have proved useful for molecular epidemiologic applications (7–19). These include the internal transcribed spacer (ITS) regions of the nuclear rRNA operon (9,10,15), the mitochondrial large subunit ribosomal RNA locus (mt LSU rRNA) (7,8,10), and the dihydropteroate synthase (DHPS) gene (10,16–19).
The ITS1 sequence is located on the nuclear rRNA operon between the genes of the 18S rRNA and the 5.8S rRNA, and ITS2 is located between the genes of the 5.8S rRNA and the 26S rRNA (22). These noncoding loci are spliced during rRNA synthesis. They show a high level of polymorphism, which has been used for genetic typing applications. The first ITS typing system was developed by Lu et al. (22). Using their nomenclature, in which ITS1 alleles are designated with an uppercase letter and ITS2 alleles with a lowercase letter, 15 ITS1 alleles (from A to O) and 14 ITS2 alleles (from a to n) have been described. Based of this amount of DNA polymorphism, a total of 210 P. jirovecii types are theoretically possible, with 59 types reported by these authors (22,23).
A second ITS typing scheme was developed by Tsolaki et al. (24–26) and is based on nucleotide variation at four positions in the ITS1 and at six positions in the ITS2. According to their nomenclature scheme, ITS1 alleles are designated by using an uppercase letter associated with a numerical subscript, and ITS2 alleles are designated by using a lowercase letter also associated with a numerical subscript. These researchers described six ITS1 alleles and nine ITS2 alleles; these numbers allow for up to 54 potential P. jirovecii ITS types, should all possible combinations exist. Laboratories currently using this typing scheme have reported ≈40 different P. jirovecii ITS types. The most frequent types are B1a3 and B2a1, which have both been identified in one third of all P. jirovecii isolates typed to date.
More recently, Nimri et al. (27) added to the count of Lee et al. by identifying 12 previously unreported ITS1 alleles and 16 previously unreported ITS2 alleles. In the study by Nimri et al., 36 ITS types were noted in 180 sequences examined from 60 samples. Although the typing methods of Lee et al. and Tsolaki et al. are not strictly identical, a general correspondence between P. jirovecii ITS types can be observed with either method. To date, approximately 87 unique ITS types have been identified by the two methods.
The amount of polymorphism reported at this locus is substantially less than that reported for ITS; nevertheless, the variation observed has helped address a number of important epidemiologic questions. The original PCR assay developed for this locus was a single-round PCR that generated a fragment of ≈360 bp (28). A nested PCR assay has also been developed (25), which has an increased sensitivity and specificity. Recently, this test was used to distinguish subclinical carriage from clinical disease (14).
Polymorphism at this locus is routinely reported at two nucleotide positions (85 and 248), showing six unique genotypes. A third variable position has been reported but is rarely seen. Mitochondrial DNA has long been accepted and used as a practical and reproducible tool to evaluate intraspecific variation. Since multiple mitochondria are present in individual organisms but display the same haplotype, mitochondrial loci are more easily detected by PCR than single-copy nuclear genes, which results in generally higher PCR sensitivities.
The DHPS locus encodes the key enzyme that is targeted by sulfonamide antimicrobial drugs. Consequently, typing efforts involving this gene have been directed primarily at demonstrating an association between treatment or prophylaxis failures and the specific mutations observed at this locus (16–19). Polymorphism at this locus has been observed primarily at amino acid positions 55 and 57, where nonsilent mutations have been shown to correlate with sulfonamide exposure and prophylaxis and treatment failure. A nested-PCR assay has been developed for this locus (29), with modifications suggested by other investigators (18). This assay is highly sensitive and specific for P. jirovecii in patients with clinical PCP. In addition to the position 55/57 mutations, nucleotide polymorphism has also been observed at several other sites in the gene (30). These mutations have not been shown to result in amino acid substitutions and have not been correlated with adverse clinical outcomes.
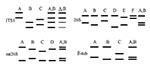
A second method for molecular typing P. jirovecii involves single-strand confirmation polymorphism (SSCP) analysis (31,32). The method consists of PCR amplification of four variable regions of P. jirovecii (Table), then detecting polymorphism by observing migration pattern variation in gel electrophoresis. The variable regions analyzed include ITS1, the intron of the nuclear 26S rRNA gene (26S), the variable region of the mt LSU rRNA, and the region surrounding an intron of the β-tubulin gene (β-tub).
A variable region amplified from a clinical specimen of a given patient with PCP can generate a simple or complex SSCP pattern. Simple patterns are made of two bands and correspond to a single allele of the genomic region. Complex patterns are made of more than two bands and have been shown to correspond to the superimposition of two simple patterns and the presence of two or, rarely, three alleles of the region (32).
According to their SSCP results, different categories of specimens can be distinguished (32). A specimen harboring a single allele at each of the four genomic regions is presumably infected with a single P. jirovecii type, and each combination of four simple SSCP patterns defines a type (Figure 1). Studies strongly suggest that a patient harboring two or more alleles of at least one of the genomic regions is coinfected with several P. jirovecii types (31). Analysis of the alleles and their abundance within the complex patterns allows identification of the coinfecting types in ≈60% of specimens coinfected with two types. Specimens producing at least one complex SSCP pattern made of three simple patterns are presumably infected with at least three types, which cannot be identified. However, the SSCP results of the latter specimens are also informative, as they often allow exclusion of certain specific types. Among 430 specimens from 15 hospitals in five European countries, three to five different simple SSCP patterns could be identified for each genomic region and 43 different P. jirovecii types. Thirty percent of the patients were infected with a single P. jirovecii, 45% with two types, and 25% with three (31,32).
A third, recently reported typing method relies on identifying the number of tandem repeats in the intron of the expression site of the major surface glycoprotein (MSG) of P. jirovecii (Table) (33). Unlike the other currently available typing methods, which rely on identifying single nucleotide polymorphisms or combinations of such polymorphisms, this method relies on characterizing the size of a region of this intron. Within this region, different P. jirovecii isolates have two to six copies of a 10-nt sequence. Typing can be performed by amplifying this region with PCR using primers flanking this region and running the PCR product on a high-resolution acrylamide gel that can separate fragments that differ in size by a few base pairs (33).
PCR followed by electrophoresis can be used to rapidly determine the number of repeats present in the intron. Because the expression site of the MSG (unlike the MSG itself) is present as a single copy per organism (34,35), a given strain of P. jirovecii will have a unique number of repeats per organism. Infections with more than one strain of P. jirovecii, which occur frequently in HIV-infected patients (20%–70% of patients) can be easily detected with this method if the different strains have a different number of repeats, since PCR amplification will result in multiple bands corresponding to the different sizes of the repeats (Figure 2).
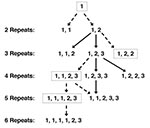
The utility of this typing method can be enhanced by sequencing the amplified PCR product because the 10-bp repeats can have one of three sequences (types 1, 2, and 3), which differ from each other by a single nucleotide. Isolates with the same number of repeats can potentially be distinguished from each other by different patterns of repeat types (e.g., three repeats of type 1, 1, 2 are different from three repeats of type 1, 2, 2). However, in isolates with multiple strains of P. jirovecii, as determined by quantifying the number of repeats, directly sequencing the PCR product will sequence only the predominant strain. Because adding or deleting 10 bases will shift the homologous sequences by 10, bases will be out of alignment downstream of the shift, making sequencing difficult. Subcloning, followed by sequencing, must be used in these circumstances to determine sequences of minority strains.
Analyzing the sequence of repeats also may provide insight into the evolution of P. jirovecii. The single base-pair changes seen in repeats likely occurred on a single occasion: since such a mutation appears to be rare (only two unique mutations have been identified) and likely does not confer a biologic advantage to the organism, the same mutation would not likely occur in the same location at different times. Thus, organisms with the 1, 1, 2 pattern of three repeats must have derived from a parental strain with a 1, 2 pattern of two repeats, rather than a 1, 1 pattern of two repeats, both of which have been seen in separate isolates (Figure 3). An analysis of multiple isolates from around the world will potentially provide information about the evolution and spread of P. jirovecii. If archival isolates can be identified and examined, we may gain additional information about the evolution of the organism over time.
Because this method was only recently described, it has been used by one group to date. In these limited studies of 147 samples from the United States and Europe, three repeats were most commonly seen, either alone or in combination (33). Two and four repeats were seen less frequently, five repeats were seen only in two mixed infections, and six repeats were seen in two isolates, one of which was in a mixed infection with three and four repeats. No pattern related to time of obtaining the sample (1974–2001) or geographic location was identified. Coinfection with more than one strain was identified in 43% of the 147 samples with this method. Additional experience with this typing method, alone and in combination with other methods, is needed to better evaluate its utility in understanding the importance of strain variation and in studying the epidemiology and biologic variability of this organism.
A potential limitation in typing that should be considered in evaluating the various approaches is the shortage of information available on marker stability and assay stability. While these data are generally lacking for most of the commonly used approaches, efforts have been made to evaluate and validate stability over time of multitarget PCR-SSCP (32). Specifically, pairs of specimens collected from the same patients during a single PCP episode were analyzed to evaluate the stability of genetic markers. Markers remained stable throughout the 8-week study, which suggests that this method was valid for most clinical applications. Less formal evaluations with direct DNA sequencing show a similar level of stability with the conserved gene loci mt LSU rRNA and DHPS (C.B. Beard, unpub. data). Unexpectedly high rates of ITS variability have been reported in samples collected from the same patient at different times during a single disease episode (36). Several possible explanations were proposed, including quantitative changes in the relative abundance of mixed P. jirovecii populations, sampling bias, intrinsic instability of the gene locus, and methodologic artifact. Others have observed similar variation patterns at this locus (C.B. Beard, unpub. data), and the explanation is a subject of debate.
Two other molecular approaches may address clinical or epidemiologic questions. These applications differ from most typing efforts in that the usual purpose of typing is to evaluate genetic polymorphism, whereas these assays examine organism numbers and viability. A recently developed quantitative PCR assay based on a conserved region of the MSG gene provides a sensitive method for quantifying organism load in oral washes of patients with suspected PCP (37,38). In the absence of a reliable culture system, PCR-based viability assays directed against mRNA targets have been also been developed (39,40). Since these applications go beyond the scope of this article, they will not be discussed further; however, they are useful for clinical and molecular epidemiologic studies.
Much consideration has been given to the question of the best genes and best approaches for molecular typing. The answer in most cases will be determined by the typing objective. One consideration is the evolutionary rate of the gene. In most eukaryotic organisms, mitochondrial DNA has been reliable for examining intraspecific variation. In P. jirovecii, the mt LSU rRNA locus has generated useful data for addressing specific epidemiologic questions (7,8,10). A greater level of intraspecific variation (22–27), however, can be detected by using the ITS locus because of its more rapid evolution. Two potential complications associated with the ITS locus are related to assay stability: the specific gene sequence, which includes a polynucleotide stretch of ≈9–12 thymidines that can lead to Taq polymerase error during amplification, and the possibility that multiple genotypes occurring in a single isolate could result from two indistinguishable sources, coinfecting P. jirovecii strains, or diploid heterozygote organisms in the sample. These concerns do not imply that the locus should not be used but only that these possibilities should be considered when interpreting the data.
The mt LSU rRNA and ITS loci are frequently used because they are not assumed to be under genetic selection and are therefore useful for elucidating molecular evolutionary phenomena that provide the basis for understanding the history of circulating strains. Sometimes, however, typing is employed specifically to determine the existence of genetic selection, such as that induced by exposure to antimicrobial agents (16–19). Care must be taken in drawing inferences from differences observed at loci that are under genetic selection, since selection can confound inferences concerning strain differences.
DNA sequencing provides the most exhaustive amount of information about any particular DNA fragment, but it is expensive and labor-intensive. Fragment analysis methods such as SSCP are simpler and less expensive, but they rely on having sequence data to characterize the patterns observed. SSCP is also limited in its ability to interpret new genotypes. Both multilocus DNA sequencing and multitarget PCR-SSCP can incorporate information from multiple genetic loci, which allows higher discriminating power to identify strain differences. Low-resolution methods (e.g., RFLP) are best used when the goal is to look only for specific RFLP-defined mutations. Either SSCP or DNA sequencing is suitable for most molecular epidemiologic studies.
The greatest need in standardization is adopting well-defined sequence types. A good example is the convention used with discussing DHPS mutations, in which the nucleotide or amino acid position is given, along with the specific identity (e.g., Thr > Ala at position 55 and Pro > Ser at position 57 to denote the DHPS double mutant genotype). Using arbitrarily defined numbers or other alphanumeric characters to define genotypes should be avoided, except perhaps for brevity in an article in which observed genotypes are all defined by nucleotide position and identity in a table. The nomenclature systems developed for the ITS locus should be reevaluated; authors’ intentions were good, but they did not account for the variation possible at that locus. A better nomenclature scheme would be to use the specific nucleotide position or variant relative to the original GenBank consensus sequence for that locus, as is typically done when reporting mutations. With the Pneumocystis genome project under way and as more genes are cataloged, nomenclature will need to be standardized further, such as with three-letter designations for genes. Within the Pneumocystis field, current practice is to use four or more letters to define some loci (e.g., DHFR, DHPS, MTLSU rRNA). Standardization will not substantially affect adopting standard typing methods and targeted gene loci, which will be of greatest use in molecular epidemiologic studies.
Dr. Beard is Chief of the Bacterial Zoonoses Branch at CDC. His research interests focus on Lyme disease, plague, tularemia, and tickborne relapsing fever. Previously, he led a research group in the Division of Parasitic Diseases at CDC that investigated the molecular epidemiology of Pneumocystis pneumonia.
References
- Vaneck J, Jirovec O, Lukes J. Interstitial plasma cell pneumonia in infants. Ann Pediatr (Paris). 1953;180:1–20.
- Edman JC, Kovacs JA, Masur H, Santi DV, Elwood HJ, Sogin ML. Ribosomal RNA sequence shows Pneumocystis carinii to be a member of the Fungi. Nature. 1988;334:519–22. DOIPubMedGoogle Scholar
- Stringer SL, Stringer JL, Blase MA, Walzer PD, Cushion MT. Pneumocystis carinii: sequence from ribosomal RNA implies a close relationship with Fungi. Exp Parasitol. 1989;68:450–61. DOIPubMedGoogle Scholar
- Stringer JR, Beard CB, Miller RF, Wakefield AE. A new name (Pneumocystis jirovecii) for pneumocystis from humans. Emerg Infect Dis. 2002;8:891–6.PubMedGoogle Scholar
- Erikson OE. Pneumocystis carinii, a parasite in lungs of mammals, referred to a new family and order (Pneumocystidaceae, Pneumocystidales, Ascomycota). Systema Ascomycetum. 1994;13:165–80.
- Latouche S, Ortona E, Mazars E, Margutti P, Tamburrini E, Siracusano A, Biodiversity of Pneumocystis carinii hominis: typing with different DNA regions. J Clin Microbiol. 1997;35:383–7.PubMedGoogle Scholar
- Wakefield AE. DNA sequences indentical to Pneumocystis carinii f. sp. carinii and Pneumocystis carinii f. sp. hominis in samples of air spora. J Clin Microbiol. 1996;34:1754–9.PubMedGoogle Scholar
- Beard CB, Carter JL, Keely SP, Huang L, Pienazek NJ, Bens MS, Genetic variation in Pneumocystis carinii isolates from different geographic regions: implications for transmission. Emerg Infect Dis. 2000;6:265–72. DOIPubMedGoogle Scholar
- Helweg-Larsen J, Tsolaki AG, Miller RF, Lundgren B, Wakefield AE. Clusters of Pneumocystis carinii pneumonia: analysis of person-to-person transmission by genotyping. QJM. 1998;91:813–20. DOIPubMedGoogle Scholar
- Miller RF, Ambrose HE, Novelli V, Wakefield AE. Probable mother-to-infant transmission of Pneumocystis carinii f. sp. hominis infection. J Clin Microbiol. 2002;40:1555–7. DOIPubMedGoogle Scholar
- Keely SP, Stringer JR, Baughman RP, Linke MJ, Walzer PD, Smulian AG. Genetic variation among Pneumocystis carinii hominis isolates in recurrent pneumocystosis. J Infect Dis. 1995;172:595–8. DOIPubMedGoogle Scholar
- Totet A, Pautard JC, Raccurt C, Roux R, Nevez G. Genotypes at the internal transcribed spacers of the nuclear rRNA operon of Pneumocystis jirovecii in non-immunosuppressed infants without severe pneumonia. J Clin Microbiol. 2003;41:1173–80. DOIPubMedGoogle Scholar
- Nevez G, Totet A, Jounieaux V, Schmit JL, Dei-Cas E, Raccurt C. Pneumocystis jirovecii internal transcribed spacer types in patients colonized by the fungus and in patients with pneumocystosis from the same French geographic region. J Clin Microbiol. 2003;41:181–6. DOIPubMedGoogle Scholar
- Wakefield AE, Lindley AR, Ambrose HE, Denis C-M, Miller RF. Limited asymptomatic carriage of Pneumocystis jirovecii in human immunodeficiency virus–infected patients. J Infect Dis. 2003;187:901–8. DOIPubMedGoogle Scholar
- Miller RF, Wakefield AE. Pneumocystis carinii genotypes and severity of pneumonia. Lancet. 1999;353:2039–40. DOIPubMedGoogle Scholar
- Helweg-Larsen J, Benfield TL, Eugen-Olsen J, Lundgren JD, Lundgren B. Effects of mutations in Pneumocystis carinii dihydropteroate synthase gene on outcome of AIDS-associated P. carinii pneumonia. Lancet. 1999;354:1347–51. DOIPubMedGoogle Scholar
- Kazanjian P, Armstrong W, Hossler PA, Burman W, Richardson J, Lee CH, Pneumocystis carinii mutations are associated with duration of sulfa or sulfone prophylaxis exposure in AIDS patients. J Infect Dis. 2000;182:551–7. DOIPubMedGoogle Scholar
- Huang L, Beard CB, Creasman J, Levy D, Duchin JS, Lee S, Sulfa or sulfone prophylaxis and geographic region predict mutations in the Pneumocystis carinii dihydropteroate synthase gene. J Infect Dis. 2000;182:1192–8. DOIPubMedGoogle Scholar
- Mei Q, Gurunathan S, Masur H, Kovacs JA. Failure of co-trimoxazole in Pneumocystis carinii infection and mutations in dihydropteroate synthase gene. Lancet. 1998;351:1631–2. DOIPubMedGoogle Scholar
- Denis CM, Mazars E, Guyot K, Odberg-Ferragut C, Viscogliosi E, Dei-Cas E, Genetic divergence at the SODA locus of six different formae speciales of Pneumocystis carinii. Med Mycol. 2000;38:289–300.PubMedGoogle Scholar
- Banerji S, Lugli EB, Miller RF, Wakefield AE. Analysis of genetic diversity at the arom locus in isolates of Pneumocystis carinii. J Eukaryot Microbiol. 1995;42:675–9. DOIPubMedGoogle Scholar
- Lu J-J, Bartlett MS, Shaw MM, Queener SF, Smith JW, Oritiz-Rivera M, Typing of Pneumocystis carinii strains that infect humans based on nucleotide sequence variations of internal transcribed spacers of rRNA genes. J Clin Microbiol. 1994;32:2904–12.PubMedGoogle Scholar
- Lee CH, Helweg-Larsen J, Tang X, Jin S, Li B, Bartlett MS, Update of Pneumocystis carinii f. sp. hominis typing based on nucleotide sequence variation in internal transcribed spacer regions of rRNA genes. J Clin Microbiol. 1998;36:734–41.PubMedGoogle Scholar
- Tsolaki AG, Miller RF, Underwood AP, Banerji S, Wakefield AE. Genetic diversity at the internal transcribed spacer regions of the rRNA operon among isolates of Pneumocystis carinii from AIDS patients with recurrent pneumonia. J Infect Dis. 1996;174:141–56. DOIPubMedGoogle Scholar
- Tsolaki AG, Beckers P, Wakefield AI. Pre-AIDS era isolates of Pneumocystis carinii f. sp. hominis: high genotype similarity with contemporary isolates. J Clin Microbiol. 1998;36:90–3.PubMedGoogle Scholar
- Tsolaki AG, Miller RF, Wakefield AE. Oropharyngeal samples for genotyping and monitoring response to treatment in AIDS patients with Pneumocystis carinii pneumonia. J Med Microbiol. 1999;48:897–905. DOIPubMedGoogle Scholar
- Nimri LF, Moura NS, Huang L, del Rio C, Rimland D, Duchin JS, Genetic diversity of Pneumocystis carinii f. sp. hominis based on variations in nucleotide sequences of the internal transcribed spacers of rRNA genes. J Clin Microbiol. 2002;40:1146–51. DOIPubMedGoogle Scholar
- Wakefield AE, Pixley FJ, Banerji S, Sinclair K, Miller RF, Moxon EF, Detection of Pneumocystis carinii with DNA amplification. Lancet. 1990;336:451–3. DOIPubMedGoogle Scholar
- Lane BR, Ast JC, Hossler PA, Mindell DP, Bartlett MS, Smith JW, Dihydropteroate synthase polymorphisms in Pneumocystis carinii. J Infect Dis. 1997;175:482–5. DOIPubMedGoogle Scholar
- Armstrong W, Meshnick S, Kazanjian P. Pneumocystis carinii mutations associated with sulfa and sulfone prophylaxis failures in immunocompromised patients. Microbes Infect. 2000;2:61–7. DOIPubMedGoogle Scholar
- Nahimana A, Blanc DS, Francioli P, Bille J, Hauser PM. Typing of Pneumocystis carinii f. sp. hominis by PCR-SSCP to indicate high frequency of co-infections. J Med Microbiol. 2000;49:753–8.PubMedGoogle Scholar
- Hauser PM, Blanc DS, Sudre P, Manoloff ES, Nahimana A, Bille J, Genetic diversity of Pneumocystis carinii in HIV-positive and -negative patients as revealed by PCR-SSCP typing. AIDS. 2001;15:461–6. DOIPubMedGoogle Scholar
- Ma L, Kutty G, Jia Q, Imamichi H, Huang L, Atzori C, Analysis of variation in tandem repeats in the intron of the major surface glycoprotein expression site of the human form of Pneumocystis carinii. J Infect Dis. 2002;186:1647–54. DOIPubMedGoogle Scholar
- Edman JC, Hatton TW, Nam M, Turner R, Mei Q, Angus CW, A single expression site with a conserved leader sequence regulates variation of expression of the Pneumocystis carinii family of major surface glycoprotein genes. DNA Cell Biol. 1996;15:989–99. DOIPubMedGoogle Scholar
- Kutty G, Ma L, Kovacs JA. Characterization of the expression site of the major surface glycoprotein of human-derived Pneumocystis carinii. Mol Microbiol. 2001;42:183–93. DOIPubMedGoogle Scholar
- Helweg-Larsen J, Lee C-H, Jin S, Hsueh JY, Benfield TL, Hansen J, Clinical correlation of variations in the internal transcribed spacer regions of rRNA genes in Pneumocystis carinii f. sp. hominis. AIDS. 2001;15:451–9. DOIPubMedGoogle Scholar
- Huang SN, Fisher SH, O’Shaughnessy E, Gill VJ, Masur H, Kovacs JA. Development of a PCR assay for diagnosis of Pneumocystis carinii pneumonia based on amplification of the multicopy major surface glycoprotein gene family. Diagn Microbiol Infect Dis. 1999;35:27–32. DOIPubMedGoogle Scholar
- Larsen HH, Masur H, Kovacs JA, Gill VJ, Silcott VA, Kogulan P, Development and evaluation of a quantitative, touch-down, real-time PCR assay for diagnosing Pneumocystis carinii pneumonia. J Clin Microbiol. 2002;40:490–4. DOIPubMedGoogle Scholar
- Maher NH, Vermund SH, Welsh DA, Dillon HK, Awooda A, Unnasch TR. Development and characterization of a molecular viability assay for Pneumocystis carinii f. sp. hominis. J Infect Dis. 2001;183:1825–7. DOIPubMedGoogle Scholar
- Latouche S, Totet A, Lacube P, Bolognini J, Nevez G, Roux P. Development of an RT-PCR on the heatshock protein 70 gene for viability detection of Pneumocystis carinii f. sp. hominis in patients with pneumocystosis and in air sample. J Eukaryot Microbiol. 2001;48(Suppl):176–7S. DOIGoogle Scholar
Figures
Table
Cite This ArticleTable of Contents – Volume 10, Number 10—October 2004
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Charles Ben Beard, CDC/NCID, Division of Vector-Borne Infectious Diseases, Rampart Rd. (Foothills Campus), Fort Collins, CO 80521, USA; fax: 970-221-6476
Top