Volume 10, Number 5—May 2004
Research
Endemic Venezuelan Equine Encephalitis in Northern Peru
Figure 3
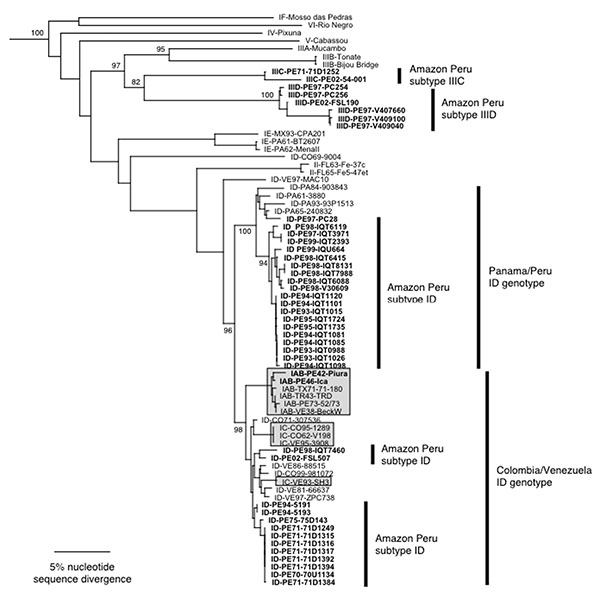
Figure 3. Phylogenetic tree of the Venezuelan equine encephalitis virus (VEEV) complex derived from partial PE2 gene sequences of Peruvian VEEV isolates and homologous sequences published previously, using the neighbor joining program implemented in PAUP 4.0 (22). The tree was rooted using an outgroup comprised of four major lineages of Eastern Equine Encephalitis virus (26). Virus strains are labeled by VEE complex subtype, abbreviated country (FL=Florida, USA) and year of isolation, followed by strain designation. Numbers indicate bootstrap values for clades to the right.
References
- Kubes V, Rios FA. The causative agent of infectious equine encephalomyelitis in Venezuela. Science. 1939;90:20–1. DOIPubMedGoogle Scholar
- Oberste MS, Fraire M, Navarro R, Zepeda C, Zarate ML, Ludwig GV, Association of Venezuelan equine encephalitis virus subtype IE with two equine epizootics in Mexico. Am J Trop Med Hyg. 1998;59:100–7.PubMedGoogle Scholar
- Lord RD. History and geographic distribution of Venezuelan equine encephalitis. Bull Pan Am Health Organ. 1974;8:100–10.PubMedGoogle Scholar
- Watts DM, Callahan J, Rossi C, Oberste MS, Roehrig JT, Wooster MT, Venezuelan equine encephalitis febrile cases among humans in the Peruvian Amazon River region. Am J Trop Med Hyg. 1998;58:35–40.PubMedGoogle Scholar
- Weaver SC, Salas R, Rico-Hesse R, Ludwig GV, Oberste MS, Boshell J, Re-emergence of epidemic Venezuelan equine encephalomyelitis in South America. VEE Study Group. Lancet. 1996;348:436–40. DOIPubMedGoogle Scholar
- Weaver SC. Recurrent emergence of Venezuelan equine encephalomyelitis. In: Scheld WM and J Hughes. Emerging infections I. Washington: ASM Press; 1998. p. 27–42.
- Johnson KM, Martin DH. Venezuelan equine encephalitis. Adv Vet Sci Comp Med. 1974;18:79–116.PubMedGoogle Scholar
- Walton TE, Grayson MA. Venezuelan equine encephalomyelitis. In: Monath TP. The arboviruses: epidemiology and ecology, vol. IV. Boca Raton (FL): CRC Press; 1988. p. 203–31.
- Oberste MS, Weaver SC, Watts DM, Smith JF. Identification and genetic analysis of Panama-genotype Venezuelan equine encephalitis virus subtype ID in Peru. Am J Trop Med Hyg. 1998;58:41–6.PubMedGoogle Scholar
- Johnson KM, Shelokov A, Peralta PH, Dammin GJ, Young NA. Recovery of Venezuelan equine encephalomyelitis virus in Panama. A fatal case in man. Am J Trop Med Hyg. 1968;17:432–40.PubMedGoogle Scholar
- Scherer WF, Madalengoitia J, Flores W, Acosta M. The first isolations of eastern encephalitis, group C, and Guama group arboviruses from the Peruvian Amazon region of western South America. Bull Pan Am Health Organ. 1975;9:19–26.PubMedGoogle Scholar
- Scherer WF, Anderson K. Antigenic and biologic characteristics of Venezuelan encephalitis virus strains including a possible new subtype isolated from the Amazon region of Peru in 1971. Am J Epidemiol. 1975;101:356–61.PubMedGoogle Scholar
- Scherer WF, Chin J. An unusual strain of Venezuelan encephalitis virus existing sympatrically with subtype I-D strains in a Peruvian rain forest. Am J Trop Med Hyg. 1983;32:871–6.PubMedGoogle Scholar
- Watts DM, Lavera V, Callahan J, Rossi C, Oberste MS, Roehrig JT, Venezuelan equine encephalitis and Oropouche virus infections among Peruvian army troops in the Amazon region of Peru. Am J Trop Med Hyg. 1997;56:661–7.PubMedGoogle Scholar
- Powers AM, Oberste MS, Brault AC, Rico-Hesse R, Schmura SM, Smith JF, Repeated emergence of epidemic/epizootic Venezuelan equine encephalitis from a single genotype of enzootic subtype ID virus. J Virol. 1997;71:6697–705.PubMedGoogle Scholar
- Wang E, Barrera R, Boshell J, Ferro C, Freier JE, Navarro JC, Genetic and phenotypic changes accompanying the emergence of epizootic subtype IC Venezuelan equine encephalitis viruses from an enzootic subtype ID progenitor. J Virol. 1999;73:4266–71.PubMedGoogle Scholar
- Weaver SC, Bellew LA, Rico-Hesse R. Phylogenetic analysis of alphaviruses in the Venezuelan equine encephalitis complex and identification of the source of epizootic viruses. Virology. 1992;191:282–90. DOIPubMedGoogle Scholar
- Weaver SC, Pfeffer M, Marriott K, Kang W, Kinney RM. Genetic evidence for the origins of Venezuelan equine encephalitis virus subtype IAB outbreaks. Am J Trop Med Hyg. 1999;60:441–8.PubMedGoogle Scholar
- Moncayo AC, Medina GM, Kalvatchev Z, Brault AC, Barrera R, Boshell J, Genetic diversity and relationships among Venezuelan equine encephalitis virus field isolates from Colombia and Venezuela. Am J Trop Med Hyg. 2001;65:738–46.PubMedGoogle Scholar
- Pfeffer M, Proebster B, Kinney RM, Kaaden OR. Genus-specific detection of alphaviruses by a semi-nested reverse transcription-polymerase chain reaction. Am J Trop Med Hyg. 1997;57:709–18.PubMedGoogle Scholar
- Black WC, Vanlandingham DL, Sweeney WP, Wasieloski LP, Calisher CH, Beaty BJ. Typing of LaCrosse, snowshoe hare, and Tahyna viruses by analyses of single-strand conformation polymorphisms of the small RNA segments. J Clin Microbiol. 1995;33:3179–82.PubMedGoogle Scholar
- Swofford DL. PAUP*. Phylogenetic analysis using parsimony (*and Other Methods). Version 4. Sunderland (MA): Sinauer Associates; 1998. p.
- Felsenstein J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution. 1985;39:783–91. DOIGoogle Scholar
- Beaty BJ, Calisher CH, Shope RE. Arboviruses. In: Schmidt NJ, RW Emmons, editors. Diagnostic procedures for viral, rickettsial and chlamydial infections. 6th ed. Washington: American Public Health Association; 1989. p. 797–855.
- Rico-Hesse R, Weaver SC, de Siger J, Medina G, Salas RA. Emergence of a new epidemic/epizootic Venezuelan equine encephalitis virus in South America. Proc Natl Acad Sci U S A. 1995;92:5278–81. DOIPubMedGoogle Scholar
- Brault AC, Powers AM, Chavez CL, Lopez RN, Cachon MF, Gutierrez LF, Genetic and antigenic diversity among eastern equine encephalitis viruses from North, Central, and South America. Am J Trop Med Hyg. 1999;61:579–86.PubMedGoogle Scholar
- Weaver SC, Dalgarno L, Frey TK, Huang HV, Kinney RM, Rice CM, Family Togaviridae. In: van Regenmortel MHV, Fauquet CM, Bishop DHL, Carstens EB, Estes MK, Lemon SM, et al. Virus taxonomy. Classification and nomenclature of viruses. Seventh report of the International Committee on Taxonomy of Viruses. San Diego: Academic Press; 2000. p. 879–89.
- Calisher CH, Shope RE, Brandt W, Casals J, Karabatsos N, Murphy FA, Proposed antigenic classification of registered arboviruses I. Togaviridae, Alphavirus. Intervirology. 1980;14:229–32. DOIPubMedGoogle Scholar
- Turell MJ, Jones JW, Sardelis MR, Dohm DJ, Coleman RE, Watts DM, Vector competence of Peruvian mosquitoes (Diptera: Culicidae) for epizootic and enzootic strains of Venezuelan equine encephalomyelitis virus. J Med Entomol. 2000;37:835–9. DOIPubMedGoogle Scholar
- Ferro C, Boshell J, Moncayo AC, Gonzalez M, Ahumada ML, Kang W, Natural enzootic vectors of Venezuelan equine encephalitis virus, Magdalena Valley, Colombia. Emerg Infect Dis. 2003;9:49–54.PubMedGoogle Scholar
- Salas RA, Garcia CZ, Liria J, Barrera R, Navarro JC, Medina G, Ecological studies of enzootic Venezuelan equine encephalitis in north-central Venezuela, 1997–1998. Am J Trop Med Hyg. 2001;64:84–92.PubMedGoogle Scholar
- Briceno Rossi AL. Rural epidemic encephalitis in Venezuela caused by a group A arbovirus (VEE). Prog Med Virol. 1967;9:176–203.PubMedGoogle Scholar
- Young NA, Johnson KM. Antigenic variants of Venezuelan equine encephalitis virus: their geographic distribution and epidemiologic significance. Am J Epidemiol. 1969;89:286–307.PubMedGoogle Scholar
- Vasconcelos PF, Travassos da Rosa JF, Travassos da Rosa AP, Degallier N, Pinheiro FP, Sa Filho GC. [Epidemiology of encephalitis caused by arbovirus in the Brazilian Amazonia]. Rev Inst Med Trop Sao Paulo. 1991;33:465–76. DOIPubMedGoogle Scholar
- Rivas F, Diaz LA, Cardenas VM, Daza E, Bruzon L, Alcala A, Epidemic Venezuelan equine encephalitis in La Guajira, Colombia, 1995. J Infect Dis. 1997;175:828–32. DOIPubMedGoogle Scholar
1Current affiliation is Instituto Venezolano de Investigaciones Científicas, Caracas, Venezuela.
2Current affiliation is Department of Biological Sciences, Ohio Northern University, Ada, Ohio.