Volume 10, Number 6—June 2004
Dispatch
Enhancing West Nile Virus Surveillance, United States
Abstract
We provide a method for constructing a county-level West Nile virus risk map to serve as an early warning system for human cases. We also demonstrate that mosquito surveillance is a more accurate predictor of human risk than monitoring dead and infected wild birds.
The introduction of West Nile virus (WNV) to the Western Hemisphere resulted in a human epidemic in New York City during 1999 (1). By 2002, WNV had spread to 44 states and the District of Columbia, with a total of 4,156 human cases of infection reported by the Centers for Disease Control and Prevention (CDC). Although a nationwide human surveillance system has been established, passive surveillance data are problematic because of variability in disease reporting. The inaccuracies in disease reporting are compounded by random variability inherent in estimating disease incidence rates, a fact that makes interpreting a risk map based on raw data difficult (2). Accounting for these issues should allow for a more precise delineation of spatial risk patterns and for improved targeting of limited prevention resources earlier in the transmission season. In addition to human cases, risk for WNV can be assessed by nonhuman surveillance systems, including infected birds and mosquitoes (3). However, these systems have not been statistically compared for their predictive ability of human risk. A quantitative assessment of the value of the nonhuman surveillance systems would also help direct resources for WNV surveillance. We provide a statistical method to estimate an accurate early assessment of human risk and to determine the predictive capabilities of nonhuman surveillance systems.
Human Surveillance Model
Human case data were taken from the weekly U.S. Geological Survey West Nile maps for the 2003 transmission season based on county-level data provided by ArboNet through voluntary reporting by state and local health officials to CDC (4). The case numbers comprise reports of mild West Nile fever as well as the more severe West Nile meningitis or encephalitis. Crude county-specific incidence rates were calculated by using the Census 2000 county population totals.
We created a human risk map for WNV based on the crude human incidence early in the transmission season, on August 13, 2003. A disease map that displays observed human incidence will show not only spatial variation in risk but also random variation resulting from low case numbers relative to the base populations. Removing random noise permits improved estimates of disease risk (2). We have approached this procedure by finding the estimates of expected incidence from a conditional autoregressive model (5,6). The model helps remove random variation based on the premise that contiguous regions tend to have similar disease risks, when compared to regions that are far apart. We applied the conditional autoregressive model to calculate expected WNV incidence rates (Appendix).
The first step was to identify the adjacent neighbors of each county by using a geographic information system (GIS, ArcView 3.2, ESRI, Redlands, CA). A data file that included the number of cases, total population, and number and names of neighboring counties for each county was then generated in SAS (SAS Institute Inc., Cary, NC). The file was imported into WinBUGS v1.4 (Imperial College, St. Mary’s, UK; and Medical Research Council, Cambridge, UK). This software implements a simulation process to estimate model parameters, including improved estimates of WNV incidence rates. These estimates were then brought back into GIS to display the human WNV risk map.
To verify the method’s potential use as an early warning system for human risk, we calculated the validity of the model-estimated risk map versus the raw incidence map from August 13 for predicting the case distribution for October 1, 2003. The two time points represent an ≈14-fold increase in total cases, from 399 to 5,685. For each of the three disease maps, counties were grouped into high- and low-risk classes on the basis of WNV incidence. High risk was defined as human incidence >1 case per 1 million population for the August 13 maps and 1 case per 100,000 for the October 1 map, findings that reflect the change in risk over time. The sensitivity of the method for predicting risk was calculated as the proportion of high-risk counties on October 1 that was correctly identified as such by the model-estimated August 13 risk map. Similarly, specificity was defined as the proportion of low-risk counties on October 1 that was correctly identified as such by the modeled risk map. The sensitivity and specificity values were compared to those obtained when the raw August 13 incidence map was used to predict risk on October 1. Measure of agreement between risk classes of the August 13 map and the October 1 map was assessed by the κ statistic, which accounts for the degree of overlap expected by chance alone; κ has a range of 0 to 1; values of <0.4 represent poor agreement (7).
Nonhuman Surveillance Model
We assessed the quantitative predictive ability of the nonhuman surveillance systems by fitting a regression model to the rate of WNV human cases for counties with the final USGS maps for the 2002 season (4). The model includes covariates for the number of virus-positive tissue samples from dead and diseased wild birds and virus-positive mosquito pools, both provided by state health officials at the county level (Appendix). Each covariate was considered together and separately to determine its contribution for predicting WNV incidence. The model was fitted by using GENMOD in SAS (SAS Institute). The contribution of nonhuman surveillance systems to variability in human risk was determined by calculating the proportion of the deviance explained (R2).
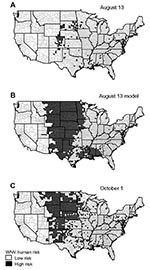
The maps of Figure 1 show the raw county-specific incidence rates for August 13, 2003 (Figure 1A), the model-estimated risk for August 13 (Figure 1B), and the raw incidence rate on October 1 (Figure 1C). The model-estimated risk surface of August 13 displays a much larger area of high risk than the reported incidence map on the same date, with 930 high-risk counties compared to 128 counties (Figure 1A and 1B). The disease map for October 1 shows a similarly larger high-risk area, with 569 counties classified as high risk (Figure 1C).
The early warning capability of our model was evaluated by comparing the validity of the raw and modeled early season disease maps for predicting the case distribution late in the transmission season (October 1). The raw data on August 13 produced a sensitivity of 19.7% (112/569) for predicting high-risk counties on October 1. In contrast, application of the model allowed for 76.1% (433/569) of the October 1 high-risk counties to be predicted, yielding a fourfold increase. This increase in sensitivity did not have a comparable negative effect on specificity, which decreased from 100% to 80.4% (2,043/2,540). In addition, the August 13 model yielded good agreement with the October 1 data, as shown by a κ statistic of 0.45 (95% confidence interval [CI] 0.42 to 0.49), whereas agreement was poor when the raw August 13 map with a κ statistic of 0.27 (CI 0.23 to 0.31) was used. Accounting for confounding caused by age distribution of WNV patients could further improve overall validity of our model.
This method has the potential to be applied in real-time to identify high-risk counties before the major influx of cases during the transmission season. The model could enable control methods to be implemented early in the season as prevention efforts before the first human case. This time advantage could provide more effective disease prevention efforts.
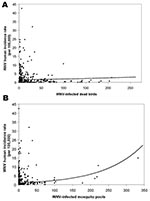
Risk modeling can also be used to effectively quantify the utility of nonhuman surveillance. Despite support for the use of bird surveillance as an early warning for WNV human risk (8–10), this system has not been statistically compared to active mosquito surveillance. The predictive ability of these surveillance systems for human risk was assessed by their inclusion as quantitative variables in a regression model. Although each variable alone was a significant predictor of human risk (χ2bird = 138.0, pbird < 0.0001; χ2mosquito = 2,605.9, pmosquito < 0.0001), the numbers of WNV-infected dead birds could only explain 2.5% of the deviance, whereas the number of WNV-positive mosquito pools explained 38%. Thus, quantitative mosquito data predict 15 times more of the variation in human cases than quantitative bird data do. The model with both covariates also explained 38% of the deviance by showing that bird data did not provide additional information about human risk (χ 2bird = 5.3, pbird=0.022; χ 2mosquito = 2,489.0, pmosquito < 0.0001). Plots of the observed and fitted incidence rates, when compared to the covariate alone, showed a much stronger positive relationship between human WNV incidence and the number of WNV-positive mosquito pools than for WNV-positive dead birds (Figure 2).
Our finding that mosquito surveillance is more sensitive to human risk than bird surveillance can be explained by the fact that human infection in the natural WNV cycle is accidental (11–14). Because birds are the zoonotic reservoir host, a WNV-infected bird only indicates enzootic transmission. For human transmission to take place, mosquito species that can act as bridge vectors must be present in sufficient numbers. Therefore, because mosquitoes represent the link to human transmission, mosquito infection prevalence should more accurately predict human risk. Furthermore, once the important human vector species can be clearly identified, the predictive ability of mosquito surveillance should increase. Standard methods for collecting mosquito data applied uniformly would also greatly aid the interpretive value of these data. Our analysis has shown that active mosquito surveillance should be emphasized in WNV surveillance systems, as it is the most sensitive marker of human risk. Surveillance systems based entirely on dead bird reports lack sensitivity for early warning as well as crucial abundance data for targeting effective prevention efforts. Entomologic surveillance should continue to be the keystone for public health programs directed toward preventing WNV infections in humans.
In summary, disease surveillance and prevention efforts could benefit from enhanced risk mapping that draws from corrected human case data and a clear understanding of the predictive ability of nonhuman surveillance.
Mr. Brownstein is completing a doctorate in the Department of Epidemiology and Public Health at Yale University School of Medicine. His research interests include the application of landscape ecology, spatial statistics, geographic information systems, and remote sensing to the surveillance of infectious diseases.
Acknowledgments
We thank Brandon Brei for his contribution.
J. S. Brownstein is supported by National Aeronautics and Space Administration Headquarters under the Earth Science Fellowship Grant NGT5-01-0000-0205 and the National Science and Engineering Research Council of Canada. This work was also supported by The Harold G. and Leila Y. Mathers Charitable Foundation (D.F.) and a U.S. Department of Agriculture-Agricultural Research Service Cooperative Agreement 58-0790-2-072 (D.F.).
References
- Lanciotti RS, Roehrig JT, Deubel V, Smith J, Parker M, Steele K, Origin of the West Nile virus responsible for an outbreak of encephalitis in the northeastern United States. Science. 1999;286:2333–7. DOIPubMedGoogle Scholar
- Mollie A. Bayesian mapping of disease. In: Gilks WR, Richardson S, Spiegelhalter DJ, editors. Markov Chain Monte Carlo in practice. New York: Chapman & Hall; 1996. p. 359–79.
- Centers for Disease Control and Prevention. Guidelines for surveillance, prevention, and control of West Nile virus infection—United States. MMWR Morb Mortal Wkly Rep. 2000;49:25–8.PubMedGoogle Scholar
- U.S. Geological Survey. West Nile virus maps—2002. Center for Integration of Natural Disaster Information; 2003 [2003 Oct 3]. Available from: http://cindi.usgs.gov/hazard/event/west_nile/west_nile.html
- Breslow N, Clayton D. Approximate inference in generalized linear mixed models. J Am Stat Assoc. 1993;88:9–25. DOIGoogle Scholar
- Clayton D, Kaldor J. Empirical Bayes estimates of age-standardized relative risks for use in disease mapping. Biometrics. 1987;43:671–81. DOIPubMedGoogle Scholar
- Landis JR, Koch GG. The measurement of observer agreement for categorical data. Biometrics. 1977;33:159–74. DOIPubMedGoogle Scholar
- Eidson M, Kramer L, Stone W, Hagiwara Y, Schmit K. Dead bird surveillance as an early warning system for West Nile virus. Emerg Infect Dis. 2001;7:631–5. DOIPubMedGoogle Scholar
- Eidson M, Komar N, Sorhage F, Nelson R, Talbot T, Mostashari F, Crow deaths as a sentinel surveillance system for West Nile virus in the northeastern United States, 1999. Emerg Infect Dis. 2001;7:615–20. DOIPubMedGoogle Scholar
- Guptill SC, Julian KG, Campbell GL, Price SD, Marfin AA. Early-season avian deaths from West Nile virus as warnings of human infection. Emerg Infect Dis. 2003;9:483–4.PubMedGoogle Scholar
- Hubalek Z, Halouzka J. West Nile fever—a reemerging mosquito-borne viral disease in Europe. Emerg Infect Dis. 1999;5:643–50. DOIPubMedGoogle Scholar
- Hulburt HS. West Nile virus infection in arthropods. Am J Trop Med Hyg. 1956;5:76–85.PubMedGoogle Scholar
- Tsai TF, Popovici F, Cernescu C, Campbell GL, Nedelcu NI. West Nile encephalitis epidemic in southeastern Romania. Lancet. 1998;352:767–71. DOIPubMedGoogle Scholar
- Savage HM, Ceianu C, Nicolescu G, Karabatsos N, Lanciotti R, Vladimirescu A, Entomologic and avian investigations of an epidemic of West Nile fever in Romania in 1996, with serologic and molecular characterization of a virus isolate from mosquitoes. Am J Trop Med Hyg. 1999;61:600–11.PubMedGoogle Scholar
Figures
Cite This ArticleTable of Contents – Volume 10, Number 6—June 2004
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Durland Fish, Department of Epidemiology and Public Health, School of Medicine, Yale University, 60 College Street, P.O. Box 208034, New Haven, CT 06520-8034, USA; fax: 203-785-3604
Top