Volume 12, Number 6—June 2006
Research
Human Parechovirus Infections in Canada
Figure 2
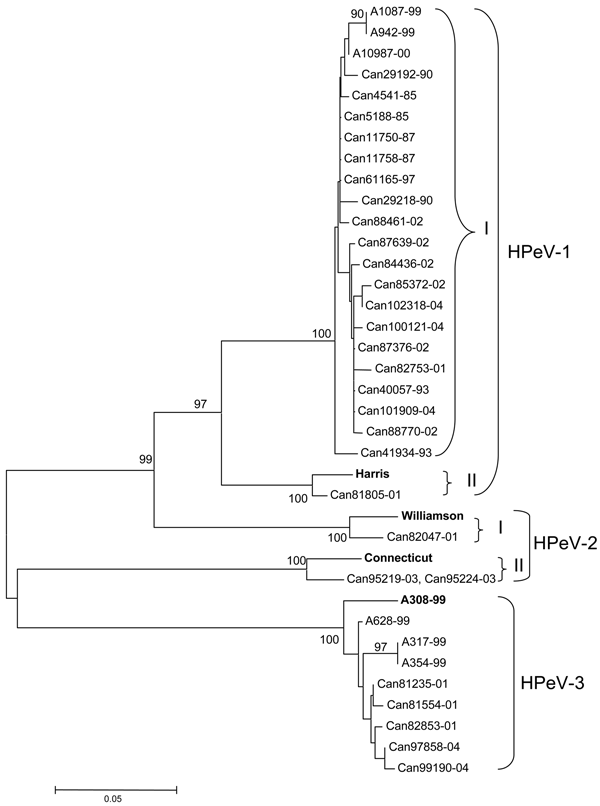
Figure 2. Phylogenetic analysis of Canadian human parechovirus (HPeV) isolates and HPeV-1 (Harris, GenBank accession no. S45208), HPeV-2 (Williamson, AJ005695, and Connecticut, AF055846), and HPeV-3 (A308-99, AB084913) reference strains based on the ClustalW alignment of the VP3 amino acid sequences. Japanese HPeV-1 (A1087-99, accession no. AB112485; A942-99, AB112486; and A10987-00, AB112487) and HPeV-3 (A628-99, accession no. AB112484; A317-99, AB112482, and A354-99, AB112483) isolates were also included in the analysis. The tree was constructed by using the neighbor-joining method with the MEGA 2 program. Bootstrap probabilities for 550 replicas are shown at the branch nodes. Only values of 70% to 100% are indicated. Boldface indicates HPeV reference strains.