Volume 13, Number 12—December 2007
Dispatch
Enhanced Subtyping Scheme for Salmonella Enteritidis
Figure 2
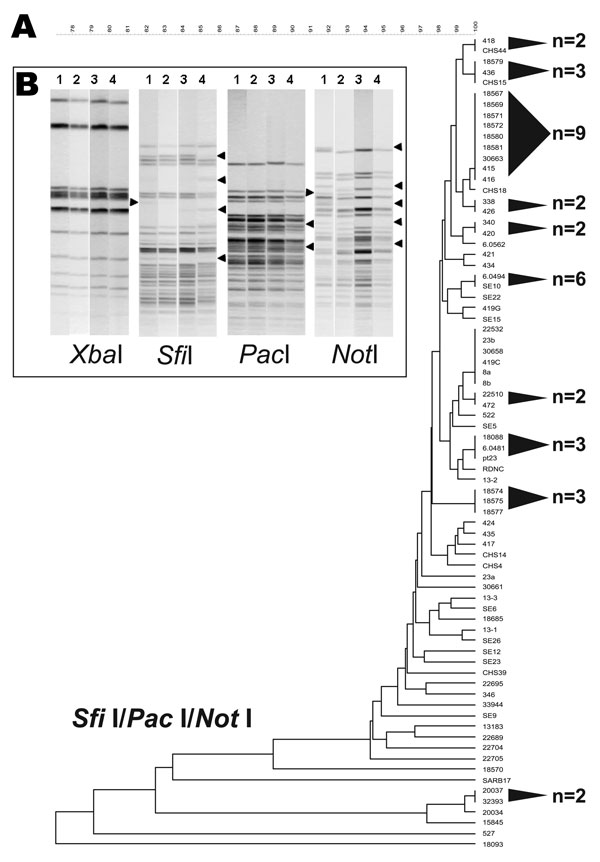
Figure 2. A 3-enzyme pulsed-field gel electrophoresis (PFGE)–based discriminatory scheme of Salmonella Enteritidis. A) Dendrogram derived from the combined analysis of PFGE data from SfiI, PacI, and NotI. Shaded cones to the right of the terminal branches denote polytomies within each dendrogram; adjacent numbers (n) show the strain totals composing their respective polytomies. A scale depicting percent divergence is presented above the dendrogram. B) Examples of S. Enteritidis strain differentiation that used SfiI, PacI, and NotI PFGE patterns. The 4 strains are numbered above the gel lanes as follows: 1, S. Enteritidis 9; 2, S. Enteritidis 12; 3, 22,704; and 4, 22,705. These strains yielded identical PFGE patterns for XbaI and BlnI. XbaI patterns shown here retain no variation among fragments. SfiI, PacI, and NotI showed examples of band polymorphism among DNA fragments.