Volume 13, Number 6—June 2007
Research
Minority-Variant pfcrt K76T Mutations and Chloroquine Resistance, Malawi
Cite This Article
Citation for Media
Abstract
Genotyping of the chloroquine-resistance biomarker pfcrt (Plasmodium falciparum chloroquine resistance transporter gene) suggests that, in the absence of chloroquine pressure, Plasmodium falciparum parasites in Malawi have reverted to chloroquine sensitivity. However, malaria infections in Africa are commonly polyclonal, and standard PCRs cannot detect minority genotypes if present in <20% of the parasites in an individual host. We have developed a multiple site–specific heteroduplex tracking assay (MSS-HTA) that can detect pfcrt 76T mutant parasites consisting of as little as 1% of the parasite population. In clinical samples, no pfcrt 76T was detected in 87 pregnant Malawian women by standard PCR. However, 22 (25%) contained minority-variant resistant genotypes detected by the MSS-HTA. These results were confirmed by subcloning and sequencing. This finding suggests that the chloroquine-resistant genotype remains common in Malawians and that PCR-undetectable drug-resistant genotypes may be present in disease-endemic populations. Surveillance for minority-variant drug-resistant mutations may be useful in making antimalarial drug policy.
Drug-resistant Plasmodium falciparum malaria continues to be a growing health problem throughout most of the world (1). To combat this threat, governments and aid agencies need accurate drug resistance surveillance data. The World Health Organization has stressed the need for methods of detecting molecular markers of drug resistance that will be useful in predicting responses to both clinical and public health interventions (2). This has been difficult in highly malaria-endemic areas, where infections are almost always polyclonal (3,4). In patients with polyclonal infections, small drug-resistant parasite populations (minority variants) may be masked by larger drug-sensitive populations because standard PCRs are relatively insensitive to minority variants (2). Therefore, new methods capable of detecting these subpopulations may lead to better drug resistance surveillance and provide a better tool to predict outcome.
We describe a new multiple site-specific heteroduplex tracking assay (MSS-HTA) for detecting the pfcrt (Plasmodium falciparum chloroquine resistance transporter gene) K76T mutation. This mutation in a putative transporter gene is well-associated with chloroquine resistance in P. falciparum (5). The MSS-HTA was compared with a standard allele-restricted PCR (ARPCR) in clinical samples from Malawi, a country where standard PCR analyses and a recent clinical trial have suggested that chloroquine-resistant malaria has disappeared (6–9).
Study Samples
Informed consent, as approved by the ethics committees of the University of North Carolina and the Malawi College of Medicine/Ministry of Health, was obtained from all participants in this research study. The Malaria and HIV in Pregnancy Study (MHP) patient samples originated from a study of pregnant women attending Queen Elizabeth Central Hospital, an urban hospital in Blantrye. The complete characteristics of the cohort and study design have been described elsewhere (10). The Mepemba and Madziabango (MM) patient samples were also collected from pregnant women as part of a pilot randomized, open-label, efficacy study of intermittent preventive treatment in pregnancy at rural health clinics. The diversity of these infections, as well as a description of the cohort and study design, has been described elsewhere (3). The characteristics of the patient samples used in this study are outlined in Table 1. All samples analyzed in this study were from filter paper blood spots of peripheral blood.
Malaria DNA Stocks
All malaria DNA used in the experiments, other than clinical samples, was from MR-4. Wild type (pfcrt K76) was from P. falciparum strain 3d7 (MR-4, MRA-102G). Two strains of mutant DNA (pfcrt 76T) were used in these experiments: P. falciparum strain K1 (MR4, MRA-159G) was used for the CVIET-resistant haplotype, based on the amino acid sequence from codon 72 to codon 76, and P. falciparum strain 7g8 (MR4, MRA-152G) was used for the SVMNT-resistant haplotype.
Generation of Heteroduplex Tracking Assay (HTA) Probe
Wild-type P. falciparum DNA was amplified with a Peltier thermal cycler (MJ Research, Waltham, MA, USA) in a volume of 50 μL. The reaction conditions consisted of 5 μL DNA, 2.5 U HotStar Taq DNA polymerase (QIAGEN, Valencia, CA, USA), 5 μL 10× PCR buffer, 1 μL deoxynucleoside triphosphate mix (catalog no. U1511, Promega, Madison, WI, USA), and 2,000 nmol/L forward and reverse primers CRT HTA F: 5′-GGAAATGGCTCACGTTTAGG-3′, and CRT HTA R: 5′-TGTGAGTTTCGGATGTTACAAAA-3′. This reaction was amplified by preheating to 95°C for 15 min, followed by 35 cycles of 95°C for 30 s, 50°C for 30 s, and 72°C for 1 min. The reaction was completed with a 10-min hold at 72°C. The 250-bp PCR product was cloned into a PCR2.1 TOPO plasmid as described in the protocol of the Topo TA cloning kit (Invitrogen, Carlsbad, CA, USA), and the sequence was confirmed at the University of North Carolina-Chapel Hill Automated DNA Sequencing Facility. Mutations were randomly introduced into the construct at the –3, –1, +1, and +3 nt relative to the single nucleotide polymorphism involved with the pfcrt K76T mutation using the QuikChange Site-Directed Mutagenesis Kit (Stratagene, La Jolla, CA, USA) (11). Potential probes were then amplified by colony PCR under the conditions noted above.
Potential probes were screened against wild-type and resistant PCR amplicons by using a heteroduplex mobility assay on an 8% polyacrylamide gel in 1% Tris-borate-EDTA buffer. After an annealing reaction, probes were evaluated for differential migration between the lanes containing the different amplicons. The annealing reaction containing 4 μL PCR product from a colony of a potential probe mixed with 4 μL PCR product of control DNA, 1 μL 100-pmol CRT HTA F primer, 1 μL 100-pmol CRT HTA R primer, and 2 μL 6× loading dye (Promega) was heated to 95°C for 2 min and then allowed to cool at 25°C for 5 min. The annealing reaction was then loaded into the wells of a nondenaturing acrylamide gel and run at a constant current of 17 mA for 5 h per gel with an SE 600 Gel Electrophoresis Unit (Amersham Biosciences, Piscataway, NJ, USA). DNA was visualized by UV after staining in an ethidium bromide solution for 20 min and destaining in double-distilled H2O for 15 min. In all, 77 potential probes were screened. The successful probe was sequenced and showed a –1 A to C mutation (GenBank accession no. submission in process).
The plasmid containing the probe was harvested by using Promega Wizard Minipreps (Promega) and then amplified according to the conditions noted above. The PCR product was the blunt-end cloned into the pT7Blue vector with the Perfectly Blunt Cloning kit (Novagen, Inc., Madison, WI, USA). The probe was radiolabeled according to the methods of Ngrenngarmlert et al. (12).
HTA
The MSS-HTA was performed under the conditions noted by Ngrenngarmlert et al. with some modifications (12). An annealing reaction consisting of 4 μL PCR product (either a control or sample DNA) was mixed with 1 μL 10× annealing buffer (1 mol/L NaCl, 100 mmol/L Tris-HCl, pH 7.5, 20 mmol/L EDTA), 2 μL 6× loading dye, 500 nmol/L CRT HTA F primer, 500 nmol/L CRT HTA R primer, and 1 μL radiolabeled probe in a total volume of 12 μL. The annealing reaction and electrophoresis were carried out under the same conditions as the heteroduplex mobility assay noted above. All MSS-HTA gels included the following controls: water, a nontemplate control PCR, and PCRs from the 3 genomic DNA stocks. The gels were dried onto filter paper (Whatman, Florham Park, NJ, USA) and exposed to BioMax MR X-ray film (Eastman Kodak, Rochester, NY, USA) for ≈48 h at 25°C. In addition, the gels were exposed to a phosphorimager screen for 48 h, and band intensities were quantified by using a GE Storm 860 Phosphorimager (Amersham Biosciences) and ImageQuant version 5.2 software (Molecular Dynamics, Sunnyvale, CA, USA).
ARPCR
ARPCR detection of pfcrt 76T was performed according to the methods described by Djimde et al. (13). The primers for the assay were modified according to Wilson et al. (8).
Minority Variant Detection
MSS-HTA and ARPCR were both run against mixtures of control DNA in quadruplicate. Differing proportions of wild-type genomic DNA and CVIET-resistant haplotype genomic DNA were mixed to a final sample concentration of 0.1 ng/μL. If a band was not visible to the eye, or only visible in 1 replicate, it was not counted.
Detection of Minor Variants in Clinical Samples
MSS-HTA was used to screen clinical samples in duplicate. All MHP samples were assayed by MSS-HTA and ARPCR. MM samples were all initially assayed by MSS-HTA. ARPCR was performed on all samples positive by HTA and on a random selection of 20 MSS-HTA–negative samples.
Ten samples that were positive by MSS-HTA were selected and Topo TA–cloned (Invitrogen). Twenty-five colonies from each of these 10 samples were screened by colony real-time PCR to determine if the plasmid construct contained a wild-type or resistant pfcrt insert (8,12). A selection of 2 mutant and 4 wild-type plasmids isolated from these colonies was then sequenced.
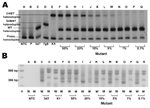
DNA from standard culture strains was used. The MSS-HTA probe formed heteroduplexes with different mobilities when annealed to P. falciparum DNA amplicons from wild-type parasites (Figure, panel A, lane C) and from parasites containing each of the 2 major resistant haplotypes: SVMNT (Figure, panel A, lane D) and CVIET (Figure, panel B, lane E).
The sensitivities of MSS-HTA and ARPCR to detect subpopulations of resistant pfcrt were tested by using artificial mixtures of wild-type and mutant genomic P. falciparum DNA. MSS-HTA detected mutant CVIET variants comprising as little as 1% of the total population (Figure, panel A, lanes N and O). In contrast, the allele-restricted PCR could not detect mutants comprising <20% of the total population (Figure, panel B, lanes I and J). In addition, the MSS-HTA accurately and reproducibly quantified mutant populations comprising as little as 1% of the sample (Table 2).
The 2 assays were then applied to clinical samples from malaria-positive Malawian pregnant women (Table 3). In total, 87 clinical samples were screened. Twenty-seven samples (MHP) were collected as part of a study of pregnant women conducted in Blantyre. CVIET-resistant haplotype P. falciparum DNA was detected in 1 sample (3.7%) by MSS-HTA and in none by ARPCR. In addition, 60 samples (MM) from 2 rural health centers were initially screened with the MSS-HTA. This method detected CVIET-resistant haplotype DNA in 21 (35%) of the clinical samples. In these samples, the amount of mutant genotype was quantified with the phosphorimager and averaged 3.3% (SD 1.4, range 1.1%–8.3%) of the parasite population. ARPCR was conducted on all samples positive by MSS-HTA as well as 20 random samples that were negative by MSS-HTA and failed to detect any samples with mutant DNA.
To confirm the presence of mutant DNA in the samples, 10 samples positive by MSS-HTA were cloned and 25 colonies from each sample were screened by real-time PCR. Of the 250 screened colonies, 6 (2.4%) had the mutant genotype in the plasmid construct. Two mutant and 4 wild-type plasmid constructs were then sequenced to confirm the MSS-HTA results. All of the mutant plasmid constructs that were sequenced contained the CVIET-resistant haplotype, and none of the wild-type plasmid constructs contained the single nucleotide polymorphism associated with pfcrt 76T.
Minority-variant drug-resistant parasite populations that were undetectable by PCR were found to be common in polyclonal Malawian P. falciparum infections. The presence of minority drug-resistant variants is consistent with results of other studies, which have shown patients with genotypicaly wild-type infections before therapy exhibiting genotypicaly mutant infections after unsuccessful chemotherapy (14,15). In Malawi, where chloroquine was replaced with sulfadoxine-pyrimethamine in 1993, the prevalence of the resistance marker pfcrt 76T, as determined by PCR, has been reported to have almost disappeared (6–9). However, our data suggest that the reversion to genotypically sensitive malaria is incomplete and that minority pfcrt 76T–bearing parasite strains are “lurking” within persons at levels undetectable by standard PCR. One caveat is that our study population comprised pregnant women with high HIV prevalence, so whether the results are applicable to the general population is unclear.
Minority-variant drug-resistant mutations are important in other diseases, such as HIV (16). The presence of minority-variant drug-resistant mutations in P. falciparum has been previously demonstrated by subcloning dihydrofolate reductase genes into yeast vectors and growing them under drug pressure (17). However, this technique cannot determine the frequency of minority variants either in a single host or in a population. To our knowledge, our results show, for the first time, that minority-variant drug-resistant mutations, representing several percentages of the parasites in a single host, are common in populations.
In response to the apparent reemergence of genotypically sensitive malaria, Laufer et al. recently completed a chloroquine efficacy trial in pediatric patients from urban Blantyre (9). The cumulative efficacy of chloroquine was 99% (95% confidence intervals 93%–100%) with only 1 treatment failure occurring in the chloroquine arm. The high efficacy rate of chloroquine therapy is not inconsistent with the results of our study. In urban Blantyre, we only found 1 patient with minority-variant pfcrt 76T. In addition, successful response to therapy requires not only susceptibility of the parasite to the drug but also factors such as acquired immunity, drug absorption, and nutrition. At this point, it is still unclear how minority-variant drug-resistant parasites will interact with these other factors. Further research in this area is needed.
Significantly more patients carried minority-variant pfcrt 76T (35%) at the rural sites (p = 0.001, Fisher exact test) than at the urban site (3.7%). Why such a marked difference was found in the prevalence of pfcrt 76T between the 2 sites is not clear. One possibility is that the transition from chloroquine to sulfadoxine-pyrimethamine may have occurred later in rural areas than in urban areas. Also, limited drug pressure may continue to be exerted on the parasites within Malawi because, as of early 2006, chloroquine was still available in local pharmacies (11). Another possible factor that may influence the prevalence of minority-variant drug-resistant parasites is external pressure from areas of high-level resistance such as Zambia and Mozambique. Mepemba and Madziabango lie on a major highway between Blantyre and the border with Zambia. Importation of cases of malaria by travel along this highway may lead to a gradient of resistance extending from the border to Blantyre. More studies on the epidemiology of minority-variant pfcrt K76T are needed to better understand the causes of this difference in the prevalence of pfcrt 76T.
In conclusion, MSS-HTAs can gather information on lurking drug resistance overlooked by standard PCRs. The method is currently performed by using radiolabeled probes, which may not be feasible in many underdeveloped countries. Implementing this method for public health purposes would require substitution of fluorescently labeled or biotinylated labeled probes for the radioisotope. MSS HTAs for other drug-resistance loci need to be developed so that the clinical and public health implications of minority variants can be fully assessed.
Dr Juliano is an Infectious Disease Fellow in training at the University of North Carolina. His research interests focus primarily on drug resistance detection in malaria from Southeast Asia and Africa. He is also interested in clinical tropical medicine.
Acknowledgment
Funding for this project comes from American Society of Public Health/Centers for Disease Control and Prevention S1935-21/21, National Institute of Allergy and Infectious Diseases (NIAID)/National Institutes of Health (NIH) AI49804, and NIH training grant DHHS/NIH/NIAID 5 T32 AI 07001-28: Training in Sexually Transmitted Diseases and AIDS. These agencies had no involvement in the design, collection, analysis, or interpretation of data in this study or in writing this article or submitting it for publication.
References
- Snow RW, Guerra CA, Noor AM, Myint HY, Hay SI. The global distribution of clinical episodes of Plasmodium falciparum malaria. Nature. 2005;434:214–7. DOIPubMedGoogle Scholar
- World Health Organization. Susceptibility of Plasmodium falciparum to antimalarial drugs. Report on global monitoring: 1996–2004. Geneva: The Organization; 2005.
- Kwiek JJ, Alker AP, Wenink EC, Chaponda M, Kalilani LV, Meshnick SR. Estimating true antimalarial efficacy by heteroduplex tracking assay in patients with complex Plasmodium falciparum infections. Antimicrob Agents Chemother. 2007;51:521–7. DOIPubMedGoogle Scholar
- Farnert A, Arez AP, Babiker HA. Genotyping of Plasmodium falciparum infections by PCR: a comparative multicentre study. Trans R Soc Trop Med Hyg. 2001;95:225–32. DOIPubMedGoogle Scholar
- Bray PG, Martin RE, Tilley L, Ward SA, Kirk K, Fidock DA. Defining the role of PfCRT in Plasmodium falciparum chloroquine resistance. Mol Microbiol. 2005;56:323–33. DOIPubMedGoogle Scholar
- Mita T, Kaneko A, Lum JK, Bwijo B, Takechi M, Zungu IL, Recovery of chloroquine sensitivity and low prevalence of the Plasmodium falciparum chloroquine resistance transporter gene mutation K76T following the discontinuance of chloroquine use in Malawi. Am J Trop Med Hyg. 2003;68:413–5.PubMedGoogle Scholar
- Kublin JG, Cortese JF, Njunju EM, Mukadam RA, Wirima JJ, Kazembe PN, Reemergence of chloroquine-sensitive Plasmodium falciparum malaria after cessation of chloroquine use in Malawi. J Infect Dis. 2003;187:1870–5. DOIPubMedGoogle Scholar
- Wilson PE, Kazadi W, Kamwendo DD, Purfield A, Meshnick SR. Prevalence of pfcrt mutations in Congolese and Malawian Plasmodium falciparum isolates as determined by a new Taqman assay. Acta Trop. 2004;93:97–106. DOIPubMedGoogle Scholar
- Laufer MK, Thesing PC, Eddington ND, Masonga R, Dzinjalamala FK, Takala SL, Return of chloroquine antimalarial efficacy in Malawi. N Engl J Med. 2006;355:1959–66. DOIPubMedGoogle Scholar
- Mwapasa V, Rogerson SJ, Kwiek JJ, Wilson PE, Milner D, Molyneux ME, Maternal syphilis infection is associated with increased risk of mother-to-child transmission of HIV in Malawi. AIDS. 2006;20:1869–77. DOIPubMedGoogle Scholar
- Resch W, Parkin N, Stuelke EL, Watkins T, Swanstrom R. A multiple-site-specific heteroduplex tracking assay as a tool for the study of viral population dynamics. Proc Natl Acad Sci U S A. 2001;98:176–81. DOIPubMedGoogle Scholar
- Ngrenngarmlert W, Kwiek JJ, Kamwendo DD, Ritola K, Swanstrom R, Wongsrichanalai C, Measuring allelic heterogeneity in Plasmodium falciparum by heteroduplex tracking assay. Am J Trop Med Hyg. 2005;72:694–701.PubMedGoogle Scholar
- Djimde A, Doumbo OK, Cortese JF, Kayentao K, Doumbo S, Diourte Y, A molecular marker for chloroquine-resistant falciparum malaria. N Engl J Med. 2001;344:257–63. DOIPubMedGoogle Scholar
- Basco LK, Ndounga M, Ngane VF, Soula G. Molecular epidemiology of malaria in Cameroon. XIV. Plasmodium falciparum chloroquine resistance transporter (PFCRT) gene sequences of isolates before and after chloroquine treatment. Am J Trop Med Hyg. 2002;67:392–5.PubMedGoogle Scholar
- Jafari S, Le Bras J, Bouchaud O, Durand R. Plasmodium falciparum clonal population dynamics during malaria treatment. J Infect Dis. 2004;189:195–203. DOIPubMedGoogle Scholar
- Kapoor A, Jones M, Shafer RW, Rhee SY, Kazanjian P, Delwart EL. Sequencing-based detection of low-frequency human immunodeficiency virus type 1 drug-resistant mutants by an RNA/DNA heteroduplex generator-tracking assay. J Virol. 2004;78:7112–23. DOIPubMedGoogle Scholar
- Mookherjee S, Howard V, Nzila-Mouanda A, Watkins W, Sibley CH. Identification and analysis of dihydrofolate reductase alleles from Plasmodium falciparum present at low frequency in polyclonal patient samples. Am J Trop Med Hyg. 1999;61:131–40.PubMedGoogle Scholar
Figure
Tables
Cite This ArticleTable of Contents – Volume 13, Number 6—June 2007
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Steven R. Meshnick, Department of Epidemiology, University of North Carolina, School of Public Health, CB#7435, Chapel Hill, NC 27599-7435, USA
Top