Volume 15, Number 6—June 2009
Letter
Crimean-Congo Hemorrhagic Fever, Southwestern Bulgaria
Figure
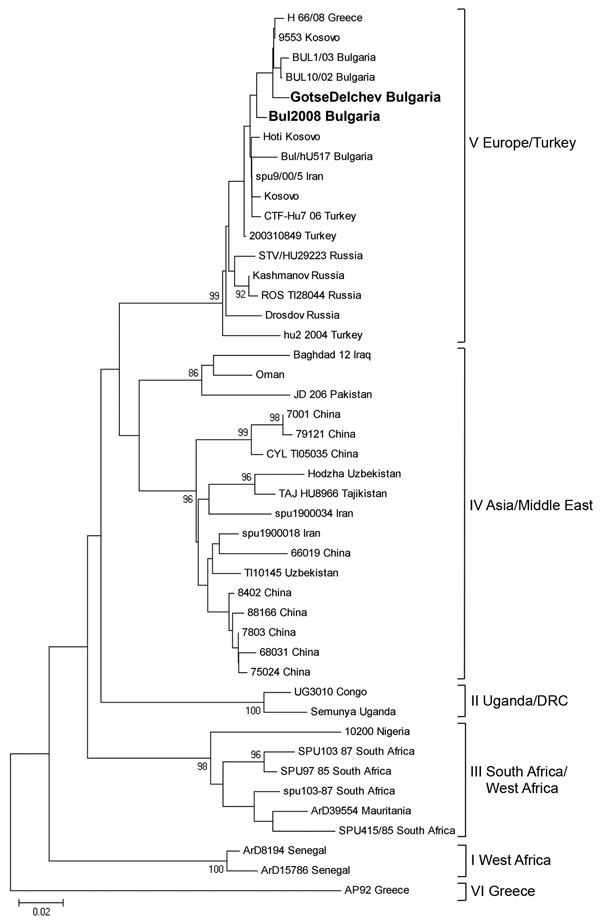
Figure. Phylogenetic tree of partial sequence (256 bp) of Crimean-Congo hemorrhagic fever (CCHF) virus nucleoprotein gene. CCHF virus sequences are listed as viral strain name and country of origin. Sequence of case 3 is designated in the tree as “Gotse Delchev Bulgaria,” and sequence of case 4 is designated as “Bul 2008 Bulgaria” (in boldface). Strain AP92, Greece, was used as an outgroup. Numbers at the nodes represent bootstrap values. Scale bar indicates number of nucleotide substitutions per site.
Page created: December 08, 2010
Page updated: December 08, 2010
Page reviewed: December 08, 2010
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.