Volume 16, Number 11—November 2010
Dispatch
Importation of Dengue Virus Type 3 to Japan from Tanzania and Côte d’Ivoire
Figure
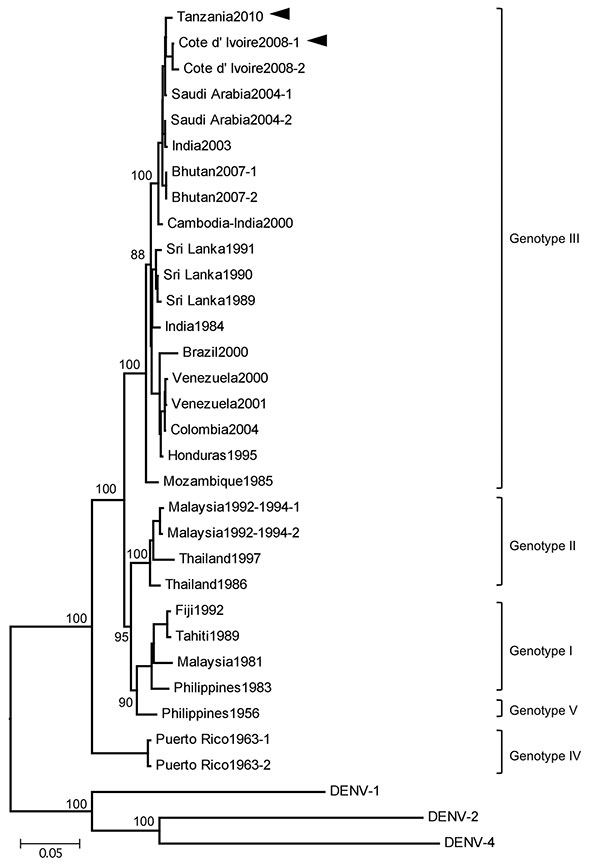
Figure. Phylogenetic tree based on the envelope genome sequence of selected dengue virus type 3 (DENV-3) strains. The tree was rooted to DENV-1, DENV-2 and DENV-4. Multiple sequence alignments were performed, and the tree was constructed by using the neighbor-joining method. The percentage of successful bootstrap replication is indicated at the nodes. DENV-3 genotypes are indicated on the right. The isolated DENV-3 strains, D3/Hu/Tanzania/NIID08/2010 strain (Tanzania2010) and D3/Hu/Côte d'Ivoire/NIID48/2008 strain (Côte d’Ivoire2008), are indicated with arrowheads. Scale bar indicates nucleotide substitutions per site.
Page created: March 09, 2011
Page updated: March 09, 2011
Page reviewed: March 09, 2011
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.