Volume 16, Number 3—March 2010
Research
New Endemic Legionella pneumophila Serogroup I Clones, Ontario, Canada
Figure 1
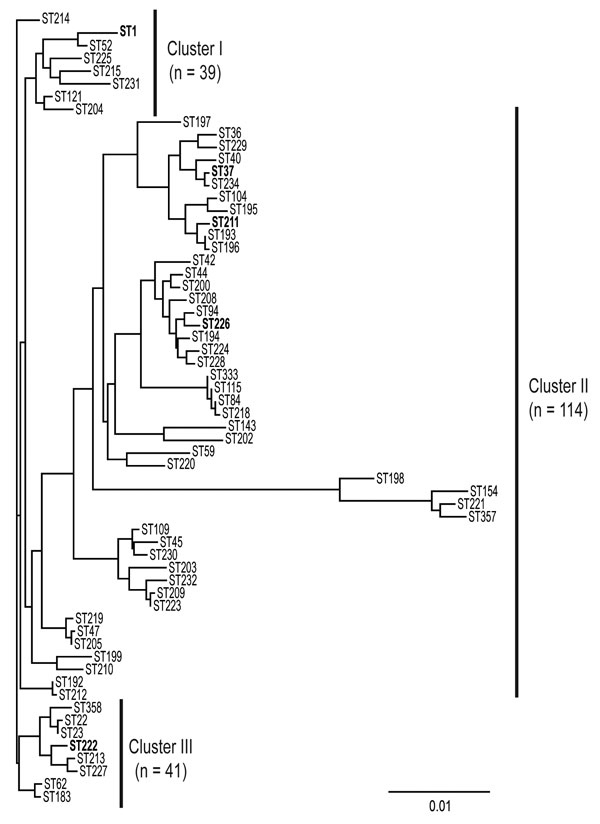
Figure 1. Phylogenetic analysis of flaA, pilE, asd, mip, mompS, proA, and neuA concatenated sequences from the 62 Legionella pneumophila serogroup 1 sequence types (STs) identified in Ontario. The tree was constructed with ClustalW2 (www.ebi.ac.uk/Tools/clustalw2/index.html) and the neighbor-joining method with 1,000 bootstrap replicates. Scale bar indicates genetic distances between sequences. STs in boldface were detected in outbreaks.
Page created: December 14, 2010
Page updated: December 14, 2010
Page reviewed: December 14, 2010
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.