Volume 16, Number 4—April 2010
Research
Escherichia albertii in Wild and Domestic Birds
Figure 2
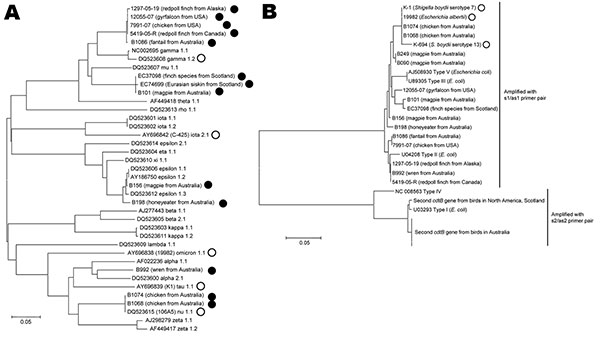
Figure 2. Neighbor-joining dendrogram based on the predicted amino acid sequences of the intimin (eae) and cytolethal distending toxin (cdtB) loci of Escherichia spp. Isolates analyzed in this study are designated by their isolate identification number; reference alleles are designated by their GenBank accession numbers. All Escherichia albertii isolates are indicated by circles or squares; all other isolates are E. coli. Scale bars indicate genetic distance. A) eae alleles carried by E. albertii isolates from birds (filled circles) represent diverse allelic subtypes and do not form separate clusters from allelic subtypes carried by isolates from humans (open circles). The eae alleles from North America and 1 from Australia (B1086) are novel but most similar to γ intimins. The eae alleles from Scotland and 1 from Australia (B101) are also novel but most similar to μ intimins. B) The E. albertii cdtB alleles from bird isolates, amplified by the s1/as1 primers, are most similar to cdtB types II, III, and V. E. albertii alleles from human isolates are marked with open circles. The bird E. albertii cdtB alleles amplified by the s2/as2 primers are distantly related to the other bird and human E. albertii alleles and are most similar to cdtB type I in E. coli. Because all the avian alleles amplified by the s2/as2 primers in each group are identical, only 1 sequence for each is shown.