Volume 16, Number 5—May 2010
Research
Anaplasma phagocytophilum from Rodents and Sheep, China
Figure 2
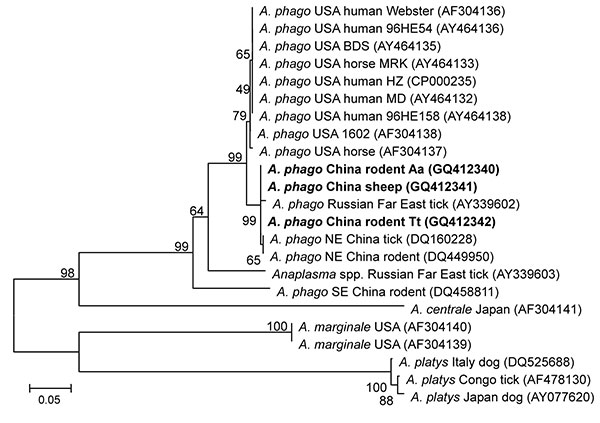
Figure 2. Phylogenetic tree based on partial (348-bp) gltA sequences of Anaplasma spp., obtained by using neighbor-joining method with Kimura 2-parameter analysis and bootstrap analysis of 1,000 replicates. Numbers on the branches indicate percentage of replicates that reproduced the topology for each clade. Parentheses enclose GenBank numbers of the sequences used in the phylogenetic analysis. Boldface indicates sequences obtained from rodents and sheep from northeastern China, May 2009. Scale bar indicates number of nucleotides per 1,000 bp. phago, phagocytophilum.
Page created: December 23, 2010
Page updated: December 23, 2010
Page reviewed: December 23, 2010
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.