Volume 16, Number 7—July 2010
Dispatch
Evolution of Seventh Cholera Pandemic and Origin of 1991 Epidemic, Latin America
Figure
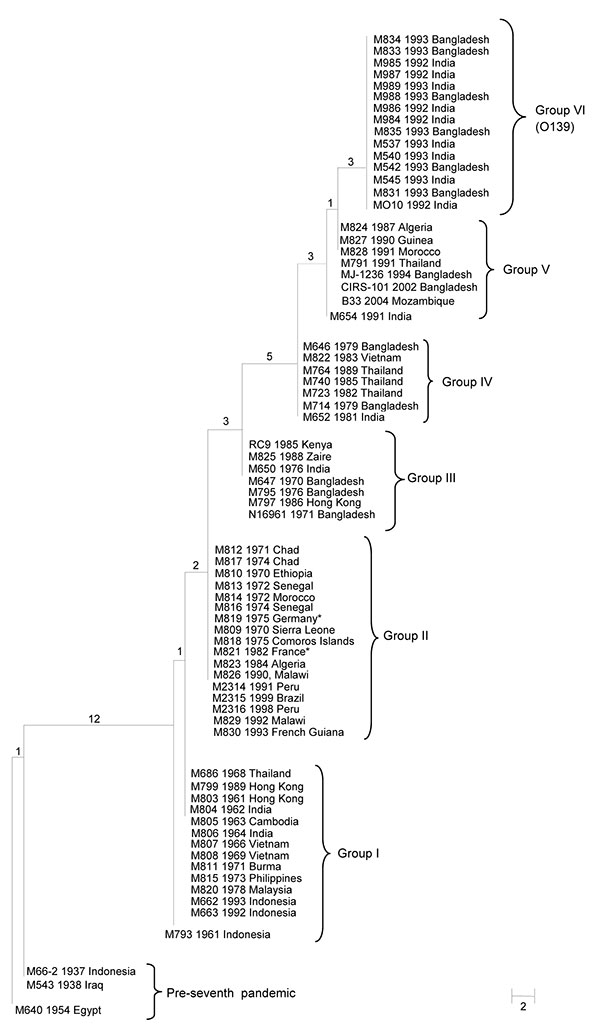
Figure. Maximum-parsimony tree of 68 seventh cholera pandemic and 3 pre–seventh cholera pandemic isolates. The tree was based on 18 N16961 seventh pandemic single-nucleotide polymorphisms (SNPs) and 12 MO10 O139 SNPs. The 3 pre–seventh pandemic isolates were used as an outgroup. Each strain name is followed by the year and location of isolation. All 15 O139 isolates had the same SNP profile and are shown as group VI. The numbers on each node represent the number of supporting SNPs. M821 and M819 from France and Germany are likely imported from either Africa or Asia. SNP data for the following isolates were obtained from GenBank: accession nos. RC9, ACHX00000000; MJ-1236, CP001385/CP001486; B33, ACHZ00000000; CIRS 101, ACVA00000000; MO10, AAKF00000000; N16961, AE003852; and M66–2, CP001233. Scale bar indicates number of nucleotide substitutions.